

Xinjiang Agricultural Sciences ›› 2023, Vol. 60 ›› Issue (7): 1766-1772.DOI: 10.6048/j.issn.1001-4330.2023.07.024
• Microbes·Animal Husbandry Veterinarian • Previous Articles Next Articles
LI Yixin1,2( ), CHEN Yong3, LIU Xiaolu2, Ainijiang Ersiman1, XU Lijuan1,4, LIU Qianqian5, BAO Xiaowei2(
), CHEN Yong3, LIU Xiaolu2, Ainijiang Ersiman1, XU Lijuan1,4, LIU Qianqian5, BAO Xiaowei2( ), SONG Suqin1(
), SONG Suqin1( )
)
Received:2022-10-07
Online:2023-07-20
Published:2023-07-11
Correspondence author:
BAO Xiaowei(1969-),female,Zhejian somgyang county, associate professor, major in food nutrition and safety,(E-mail)408531623@qq.com;Supported by:
李怡歆1,2( ), 陈勇3, 刘晓禄2, 艾尼江·尔斯满1, 徐李娟1,4, 刘倩倩5, 包晓玮2(
), 陈勇3, 刘晓禄2, 艾尼江·尔斯满1, 徐李娟1,4, 刘倩倩5, 包晓玮2( ), 宋素琴1(
), 宋素琴1( )
)
通讯作者:
包晓玮(1969-),女,浙江松阳人,副教授,博士,研究方向为食品营养与安全,(E-mail)408531623@qq.com;作者简介:李怡歆(1997-),女,新疆伊犁人,硕士研究生,研究方向为放线菌活性,(E-mail)2463514716@qq.com
基金资助:CLC Number:
LI Yixin, CHEN Yong, LIU Xiaolu, Ainijiang Ersiman, XU Lijuan, LIU Qianqian, BAO Xiaowei, SONG Suqin. Isolation, identification and activity of halophilic bacteria from Dabancheng Salt Lake, Xinjiang[J]. Xinjiang Agricultural Sciences, 2023, 60(7): 1766-1772.
李怡歆, 陈勇, 刘晓禄, 艾尼江·尔斯满, 徐李娟, 刘倩倩, 包晓玮, 宋素琴. 新疆达坂城盐湖嗜盐细菌分离鉴定及活性分析[J]. 新疆农业科学, 2023, 60(7): 1766-1772.
Add to citation manager EndNote|Ris|BibTeX
URL: http://www.xjnykx.com/EN/10.6048/j.issn.1001-4330.2023.07.024
| 培养基 Culture medium | 嗜盐细菌属 Halophilic bacteria | 嗜盐细菌种 Halophilic bacterial species |
|---|---|---|
| M-2(海藻糖-脯氨酸) | 类芽孢杆菌属(Paenibacillus) | Paenibacillus peoriae |
| MM琼脂 | 代谢杆菌(Metabacillus) | Metabacillus sediminilitoris |
| M-7(甘油-天门冬酰胺) | 类诺卡氏菌属(Nocardioides) 动性球菌属(Planococcus) 冢村氏菌属(Tsukamurella) 短杆菌属(Brevibacterium) 假黄色单胞菌属(Pseudoxanthomonas) 链霉菌属(Streptomyces) 红球菌属(Rhodococcus) 小单胞菌属(Micromonospora) 芽孢杆菌属(Bacillus) 细杆菌属(Microbacterium paraoxydans) 微球菌属(Micrococcus) 拟诺卡氏菌属(Nocardiopsis) | Nocardioides furvisabuli、Nocardioides LTZG-s、Tsukamurella conjunctivitidis、Brevibacterium epidermidis、Pseudoxanthomonas japonensis、Streptomyces longispororuber、Rhodococcus qingshengii、Micromonospora oryzae、Bacillus paramycoides、Microbacterium paraoxydans、Micrococcus antarcticus、Tsukamurella_sinensis、Nocardiopsis dassonvillei |
| M-1(TWYE) | 链霉菌属(Streptomyces) | Streptomyces pratensis |
Tab.1 Species of halophilic bacteria isolated from different media
| 培养基 Culture medium | 嗜盐细菌属 Halophilic bacteria | 嗜盐细菌种 Halophilic bacterial species |
|---|---|---|
| M-2(海藻糖-脯氨酸) | 类芽孢杆菌属(Paenibacillus) | Paenibacillus peoriae |
| MM琼脂 | 代谢杆菌(Metabacillus) | Metabacillus sediminilitoris |
| M-7(甘油-天门冬酰胺) | 类诺卡氏菌属(Nocardioides) 动性球菌属(Planococcus) 冢村氏菌属(Tsukamurella) 短杆菌属(Brevibacterium) 假黄色单胞菌属(Pseudoxanthomonas) 链霉菌属(Streptomyces) 红球菌属(Rhodococcus) 小单胞菌属(Micromonospora) 芽孢杆菌属(Bacillus) 细杆菌属(Microbacterium paraoxydans) 微球菌属(Micrococcus) 拟诺卡氏菌属(Nocardiopsis) | Nocardioides furvisabuli、Nocardioides LTZG-s、Tsukamurella conjunctivitidis、Brevibacterium epidermidis、Pseudoxanthomonas japonensis、Streptomyces longispororuber、Rhodococcus qingshengii、Micromonospora oryzae、Bacillus paramycoides、Microbacterium paraoxydans、Micrococcus antarcticus、Tsukamurella_sinensis、Nocardiopsis dassonvillei |
| M-1(TWYE) | 链霉菌属(Streptomyces) | Streptomyces pratensis |
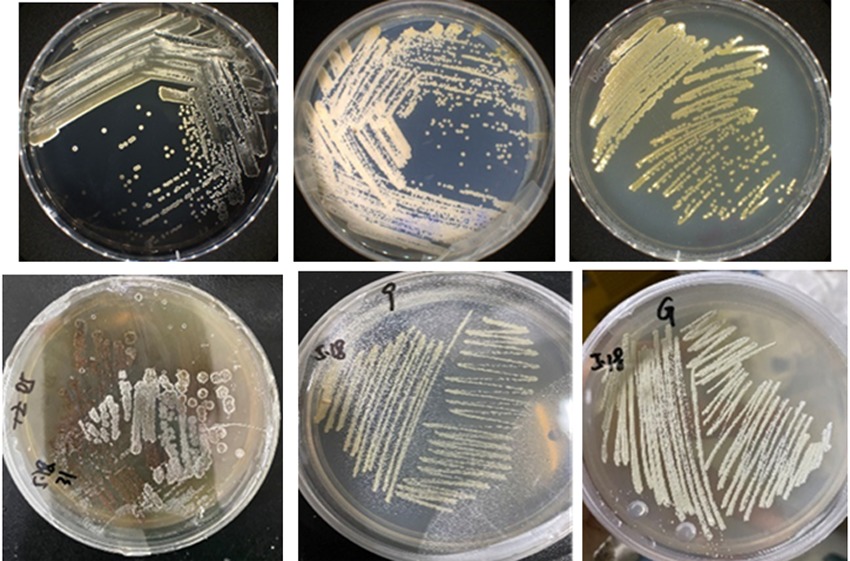
Fig.1 Part of halophilic bacteria in Dabancheng Salt Lake, Xinjiang Note:a:Strain DBC23; b:Strain DBC30; c:Strain DBC31; d:Strain DBC5; e:Strain DBC9; f:Strain DBC10
| 序列号 NO. | 菌株编号 NO. | 最近相似菌株名称 The name of most similarity bacteria | 相似度 Similarity | 收藏号 Accession NO. |
|---|---|---|---|---|
| 1 | DBC1 | Bacillus mobilis | 99.51% | MACF01000036 |
| 2 | DBC2 | Pseudoxanthomonas japonensis | 99.14% | AB008507 |
| 3 | DBC3 | Micromonospora oryzae | 99.35% | AB981052 |
| 4 | DBC4 | Rhodococcus qingshengii | 99.64% | LRRJ01000016 |
| 5 | DBC5 | Streptomyces longispororuber | 99.71% | AB184440 |
| 6 | DBC7 | Brevibacterium epidermidis | 99.60% | BCSJ01000023 |
| 7 | DBC9 | Tsukamurella conjunctivitidis | 99.71% | MK583684 |
| 8 | DBC25 | LTZG_s | 99.10% | LTZG01000007 |
| 9 | DBC23 | Nocardioides alpinus | 97.43% | GU784866 |
| 10 | DBC10 | Streptomyces pratensis | 99.85% | JQ806215 |
| 11 | DBC12 | Metabacillus sediminilitoris | 98.20% | MN067806 |
| 12 | DBC13 | MRTF_s | 98.90% | MRTF01000019 |
| 13 | DBC18 | Bacillus paramycoides | 95.93% | MAOI01000012 |
| 14 | DBC22 | Microbacterium paraoxydans | 99.22% | BCRH01000180 |
| 15 | DBC24 | Micrococcus antarcticus | 98.93% | AJ005932 |
| 16 | DBC26 | Tsukamurella sinensis | 99.64% | KP888571 |
| 17 | DBC30 | Nocardiopsis dassonvillei | 100% | ABUI01000017 |
| 18 | DBC31 | Nocardiopsis crassaminis | 99.82% | LR606207 |
Tab.2 Sequence similarity of 16S rRNA between isolated strains and known strains
| 序列号 NO. | 菌株编号 NO. | 最近相似菌株名称 The name of most similarity bacteria | 相似度 Similarity | 收藏号 Accession NO. |
|---|---|---|---|---|
| 1 | DBC1 | Bacillus mobilis | 99.51% | MACF01000036 |
| 2 | DBC2 | Pseudoxanthomonas japonensis | 99.14% | AB008507 |
| 3 | DBC3 | Micromonospora oryzae | 99.35% | AB981052 |
| 4 | DBC4 | Rhodococcus qingshengii | 99.64% | LRRJ01000016 |
| 5 | DBC5 | Streptomyces longispororuber | 99.71% | AB184440 |
| 6 | DBC7 | Brevibacterium epidermidis | 99.60% | BCSJ01000023 |
| 7 | DBC9 | Tsukamurella conjunctivitidis | 99.71% | MK583684 |
| 8 | DBC25 | LTZG_s | 99.10% | LTZG01000007 |
| 9 | DBC23 | Nocardioides alpinus | 97.43% | GU784866 |
| 10 | DBC10 | Streptomyces pratensis | 99.85% | JQ806215 |
| 11 | DBC12 | Metabacillus sediminilitoris | 98.20% | MN067806 |
| 12 | DBC13 | MRTF_s | 98.90% | MRTF01000019 |
| 13 | DBC18 | Bacillus paramycoides | 95.93% | MAOI01000012 |
| 14 | DBC22 | Microbacterium paraoxydans | 99.22% | BCRH01000180 |
| 15 | DBC24 | Micrococcus antarcticus | 98.93% | AJ005932 |
| 16 | DBC26 | Tsukamurella sinensis | 99.64% | KP888571 |
| 17 | DBC30 | Nocardiopsis dassonvillei | 100% | ABUI01000017 |
| 18 | DBC31 | Nocardiopsis crassaminis | 99.82% | LR606207 |
| 植物病原菌Plant pathogen | |||||
|---|---|---|---|---|---|
| 嗜盐放线菌 Halophilic actinomycetes | 南瓜枯萎病菌 F.oxysporum | 水稻恶苗病菌 F.moliniforme | 棉花枯萎病菌 F.oxysporum | 核桃腐烂病菌 Cytcospora sp. | 苹果树腐烂病菌 Valsa mali var.mali |
| DBC5 | — | + | — | — | — |
| DBC23 | — | — | — | — | — |
| DBC10 | — | — | — | — | — |
| DBC26 | — | — | + | — | — |
| DBC30 | — | — | — | — | — |
| DBC31 | — | — | + | — | — |
Tab.3 Antibacterial activity of halophilic actinomycetes in Dabancheng Salt Lake
| 植物病原菌Plant pathogen | |||||
|---|---|---|---|---|---|
| 嗜盐放线菌 Halophilic actinomycetes | 南瓜枯萎病菌 F.oxysporum | 水稻恶苗病菌 F.moliniforme | 棉花枯萎病菌 F.oxysporum | 核桃腐烂病菌 Cytcospora sp. | 苹果树腐烂病菌 Valsa mali var.mali |
| DBC5 | — | + | — | — | — |
| DBC23 | — | — | — | — | — |
| DBC10 | — | — | — | — | — |
| DBC26 | — | — | + | — | — |
| DBC30 | — | — | — | — | — |
| DBC31 | — | — | + | — | — |
| 菌株编号 Strain No. | 抑制率 Inhibition rate |
|---|---|
| DBC5 | 96.4% |
| DBC10 | 87.8% |
| DBC9 | 91.8% |
| DBC3 | NA |
| DBC26 | 83.2% |
| DBC24 | 16.7% |
Tab.4 Inhibition of crude extract of salt lake actinomycetes fermentation broth on HepG2 cells
| 菌株编号 Strain No. | 抑制率 Inhibition rate |
|---|---|
| DBC5 | 96.4% |
| DBC10 | 87.8% |
| DBC9 | 91.8% |
| DBC3 | NA |
| DBC26 | 83.2% |
| DBC24 | 16.7% |
| [1] | 高波, 吉兰泰盐湖土壤中嗜盐碱放线菌的分离鉴定及四氢嘧啶工程菌株的构建[D] 杨凌: 西北农林科技大学, 2016. |
| GAO Bo. Isolation and identification of saline-alkali actinomycetes from soil of Jilantai Salt Lake and construction of tetrahydropyrimidine engineering strain[D]. Yangling: Northwest A&F University, 2016. | |
| [2] |
Hacěne H, Bhatnagar J C, Baratti B. Biodiversity of prokaryotic microflora in El Golea Salt lake, Algerian Sahara[J]. Journal of Arid Environments, 2004, 58(3):273-284.
DOI URL |
| [3] |
Shantikumar L. Author's personal copy Optimisation of process parameters for growth and bioactive metabolite produced by a salt-tolerant and alkaliphilic actinomycete, Streptomyces tanashiensis strain A2D[J]. Journal de Mycologie Médicale, 2009, 19(4):225-233.
DOI URL |
| [4] | 吴海平, 王真辉, 杨礼富. 新疆达坂盐湖沉积土壤嗜盐细菌的定向富集与多样性分析[J]. 微生物学通报, 2010, 37(7):956-961. |
| WU Haiping, WANG Zhenhui, YANG Lifu. Analysis on the diversity and enrichment of halophilic bacteria in sediments of daban salt lake, Xinjiang[J]. Microbiology Bulletin, 2010, 37(7):956-961. | |
| [5] | 刘会强, 赵春梅, 黄瑞虎. 新疆达坂城盐湖中度嗜盐菌的分类研究[J]. 新疆师范大学学报(自然科学版), 2007,(3):146-149. |
| LIU Huiqiang, ZHAO Chunmei, HUANG Ruihu. Classification of moderate halophilic bacteria in Dabancheng Salt Lake, Xinjiang[J]. Journal of Xinjiang Normal University (Natural Science Ed.), 2007,(3):146-149. | |
| [6] | 张立丰. 新疆达坂城盐湖嗜盐古菌16S rDNA序列分析和细菌视紫红质基因序列的研究[D]. 乌鲁木齐: 新疆师范大学, 2006. |
| ZHANG Lifeng. 16S rDNA sequence analysis and bacterial rhodopsin gene sequence of halophilic archaea from Dabancheng Salt Lake, Xinjiang[D]. Urumqi: Xinjiang Normal University, 2006. | |
| [7] | 关统伟. 新疆罗布泊盐湖放线菌多样性及多相分类[D]. 成都: 四川农业大学, 2012. |
| GUAN Tongwei. Diversity and multiphase classification of actinomycetes in Lop Nur Salt Lake, Xinjiang[D]. Chengdu: Sichuan Agricultural University, 2012. | |
| [8] | 宋阳, 王秋菊, 杨旭, 等. 新疆盐碱土壤耐盐碱放线菌筛选初步鉴定[J]. 生物化工, 2020, 6(3):80-84. |
| SONG Yang, WANG Qiuju, YANG Xu, et al. Preliminary screening and identification of saline-alkali tolerant actinomycetes in xinjiang saline-alkali soil[J]. Biochemical Engineering, 2020, 6(3):80-84. | |
| [9] | 李文均, 焦建宇. 放线菌分类地位的变迁及其系统学研究最新进展[J]. 微生物学杂志, 2020, 40(1):1-14. |
| LI Wenjun, JIAO Jainyu. Changes in taxonomic status of actinomycetes and recent advances in systematic research[J]. Journal of Microbiology, 2020, 40(1):1-14. | |
| [10] |
Skehan P, Storeng R, Scudiero D, et al. New colorimetric cytotoxicity assay for anticancer-drug screening[J]. Journal of the National Cancer Institute, 1990, 82(13):1107-1112.
DOI PMID |
| [11] | 任海柯, 死海极端高盐环境放线菌资源初步研究[D]. 杨凌: 西北农林科技大学, 2012. |
| REN Haike. Preliminary study on actinomycetes resources in extreme saline environment of the Dead Sea[D]. Yangling: Northwest A&F University, 2012. | |
| [12] | 李二阳, 马雪莉, 吕杰, 等. 新疆天山北坡不同盐湖微生物菌群结构及其影响因子[J]. 生态学报, 2021, 41(18):7212-7225. |
| LI Eryang, MA Xueli, LV Jie, et al. Microflora structure and its influencing factors in different salt lakes on the Northern Slope of Tianshan Mountains, Xinjiang[J]. Acta Ecologica Sinica, 2021, 41(18):7212-7225. | |
| [13] | 唐蜀昆, 李文均, 张永光, 等. 嗜盐放线菌生物学特性初步研究[J]. 微生物学通报, 2003,(4):15-19. |
| TANG Shukun, LI Wenjun, ZHANG Yonghua, et al. Preliminary study on biological characteristics of halophilic actinomycetes[J]. Microbiology Bulletin, 2003,(4):15-19. | |
| [14] |
Babich H, Stotzky G. Temperature, pH, salinity, hardness, and particulates mediate nickel toxicity to eubacteria, an actinomycete, and yeasts in lake, simulated estuarine, and sea waters[J]. Aquatic Toxicology, 1983, 3(3):195-208.
DOI URL |
| [1] | WANG Dandan, LI Yan, ZHANG Qingyin, Li Shidong, PANG Yongchao, MA Kunzhi, MA Long, NIU Ruisheng, ZHONG Zengming, QI Lianfen, SHI Jianhua. Effects of different microbial treatments on tomato soil microbial diversity [J]. Xinjiang Agricultural Sciences, 2023, 60(9): 2248-2257. |
| [2] | ZHOU Shijie, DONG Yiqiang, Asitaiken Julihaiti, NIE Tingting, JIANG Anjing, AN Shazhou. Quantitative characteristics and diversity of sagebrush desert plant communities on the northern slope of Tianshan Mountains [J]. Xinjiang Agricultural Sciences, 2023, 60(9): 2298-2305. |
| [3] | LI Kailiang, ZHANG Zhenyu, HU Hongying. Structure and diversity analysis of insect community in jujube orchard of Hami Area in Xinjiang [J]. Xinjiang Agricultural Sciences, 2023, 60(8): 2028-2037. |
| [4] | CHEN Yihuang, XING Li, DONG Zhenzhen, MA Xiaomei, HUANG Jianjun, LUO Xiaoxia. Isolation and antagonistic activity screening of Actinomycetes from rhizosphere soil of Tamarix [J]. Xinjiang Agricultural Sciences, 2023, 60(7): 1790-1797. |
| [5] | CHEN Kaixu, GUO Cuijie, YANG Fan, REN Feier, LI Xiaobin, LIU Wujun. Genetic diversity analysis of xinjiang sheep with fine wool based on whole-genome Re-sequencing [J]. Xinjiang Agricultural Sciences, 2023, 60(5): 1292-1300. |
| [6] | LIU Shengxue, LI Siqi, WANG Xiaodong, YANG Desong. Diversity analysis of dsRNA in Alternaria solani [J]. Xinjiang Agricultural Sciences, 2023, 60(12): 3057-3064. |
| [7] | YUE Li, WANG Hui, Shanqimike , Zaituniguli Kuerban, TU Zhendong. Microbial diversity analysis of sweet sorghum silage using High-Throughput sequencing [J]. Xinjiang Agricultural Sciences, 2023, 60(11): 2742-2750. |
| [8] | LUO Ying, Nurziya Yalimaimaiti, JIA Wenjie, LIU Jieying, JIA Peisong. Identification and genetic diversity of wild Macrolepiota in Xinjiang [J]. Xinjiang Agricultural Sciences, 2023, 60(10): 2501-2508. |
| [9] | LI Feng, LAI Ganggang, ZHAO Zhihui, CHEN Xiaofei, ZHU Tiansheng. Identification and 16S rRNA gene diversity analysis of salt cedar(Tamarix chinensis)witches'-broom [J]. Xinjiang Agricultural Sciences, 2023, 60(10): 2551-2557. |
| [10] | YAO Yingying, LI Jiahui, LI Haiying, WU Yingping, ZHAO Xiaoyu, ZHOU Jun, ZHAO Quanzhuang, LI Zongfu. Analysis of genetic diversity of Yemili chicken by microsatellite [J]. Xinjiang Agricultural Sciences, 2023, 60(10): 2574-2582. |
| [11] | ZHAO Shuangyin, WANG Weiran, YAN Xuexue, Bimairemu Abuduaihaiti, DONG Jie, Tuerxon Tuerhong, Alip Aierxi. Analysis of Quality Traits and Genetic Diversity of 120 Cotton Germplasm Resources [J]. Xinjiang Agricultural Sciences, 2023, 60(1): 17-24. |
| [12] | LI Yueyan, LI Yushan, WANG Fan, GUO Yawen, WANG Feiyan, GAO Jie, SONG Yu. Genetic Diversity and Cluster Analysis of Tomato Fruit Characters in Different Varieties [J]. Xinjiang Agricultural Sciences, 2022, 59(9): 2147-2157. |
| [13] | LI Xuanwen, XIONG Zhi, WANG Jinhua, ZHOU Yiping, XIONG Zhongping. Study on the Diversity of Gut Bacteria from Adults of Dendrolimus kikuchii [J]. Xinjiang Agricultural Sciences, 2022, 59(9): 2276-2287. |
| [14] | YUAN Lei, JI Xuehua, ZHANG Guoru, SHI Linyuan, GUO Heyao, TANG Yaping, YANG Tao, YANG Shengbao. Genetic Diversity and Cluster Analysis on the Main Fruit Characters of 52 Accessions of Capsicum [J]. Xinjiang Agricultural Sciences, 2022, 59(8): 1935-1944. |
| [15] | WU Qiaoyu, HE Tianjiu. Morphological and ISSR Analysis of Purple Sweet Potato Resources [J]. Xinjiang Agricultural Sciences, 2022, 59(7): 1625-1631. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||