

Xinjiang Agricultural Sciences ›› 2023, Vol. 60 ›› Issue (11): 2627-2637.DOI: 10.6048/j.issn.1001-4330.2023.11.004
• Crop Genetics and Breeding · Germplasm Resources·Molecular Genetics·Soil Fertilizer • Previous Articles Next Articles
HU Wenran( ), ZHAO Zhun, SHAO Wukui, HUANG Quansheng(
), ZHAO Zhun, SHAO Wukui, HUANG Quansheng( )
)
Received:2023-02-11
Online:2023-11-20
Published:2023-12-07
Correspondence author:
Huang Quansheng (1964-), Male, Urumqi, researcher, doctor, research direction: molecular biology of crop resistance, (E-mail) hquansheng@126.com
Supported by:通讯作者:
黄全生(1964-),男,新疆人,研究员,博士,研究方向为农作物生物技术,(E-mail)hquansheng@126.com
作者简介:胡文冉(1974-),女,河南人,研究员,博士,研究方向为棉花生物技术,(E-mail)huwran@126.com
基金资助:CLC Number:
HU Wenran, ZHAO Zhun, SHAO Wukui, HUANG Quansheng. Identification of TCP family and analysis of tissue expression in upland cotton[J]. Xinjiang Agricultural Sciences, 2023, 60(11): 2627-2637.
胡文冉, 赵准, 邵武奎, 黄全生. 陆地棉TCP家族基因鉴定及组织表达分析[J]. 新疆农业科学, 2023, 60(11): 2627-2637.
Add to citation manager EndNote|Ris|BibTeX
URL: http://www.xjnykx.com/EN/10.6048/j.issn.1001-4330.2023.11.004
| 蛋白质类型 Type of protein | 亚家族 Subfamilies | 拟南芥 Arabido- psis | 水稻 Rice | 陆地棉 Upland cotton |
|---|---|---|---|---|
| Class I | 13 | 10 | 39 | |
| Class Ⅱ | CYC/TB1 | 3 | 3 | 7 |
| CIN | 8 | 9 | 17 |
Tab.1 Comparison of TCP proteins number in Arabidopsis, rice and upland cotton
| 蛋白质类型 Type of protein | 亚家族 Subfamilies | 拟南芥 Arabido- psis | 水稻 Rice | 陆地棉 Upland cotton |
|---|---|---|---|---|
| Class I | 13 | 10 | 39 | |
| Class Ⅱ | CYC/TB1 | 3 | 3 | 7 |
| CIN | 8 | 9 | 17 |
| 基因类型 Gene type | 外显子 数量 Number of exons | 基因数量 Number of gene | 蛋白质大小 Protein length(aa) | |
|---|---|---|---|---|
| Class I | 1 | 38 | 150-550 | |
| 2 | 1 | 353 | ||
| Class Ⅱ | CYC/TB1 | 1 | 3 | 367-501 |
| 2 | 4 | 325-414 | ||
| CIN | 1 | 14 | 285-463 | |
| 2 | 3 | 266-451 | ||
Tab.2 The number of exons and protein length of TCP genes in upland cotton
| 基因类型 Gene type | 外显子 数量 Number of exons | 基因数量 Number of gene | 蛋白质大小 Protein length(aa) | |
|---|---|---|---|---|
| Class I | 1 | 38 | 150-550 | |
| 2 | 1 | 353 | ||
| Class Ⅱ | CYC/TB1 | 1 | 3 | 367-501 |
| 2 | 4 | 325-414 | ||
| CIN | 1 | 14 | 285-463 | |
| 2 | 3 | 266-451 | ||
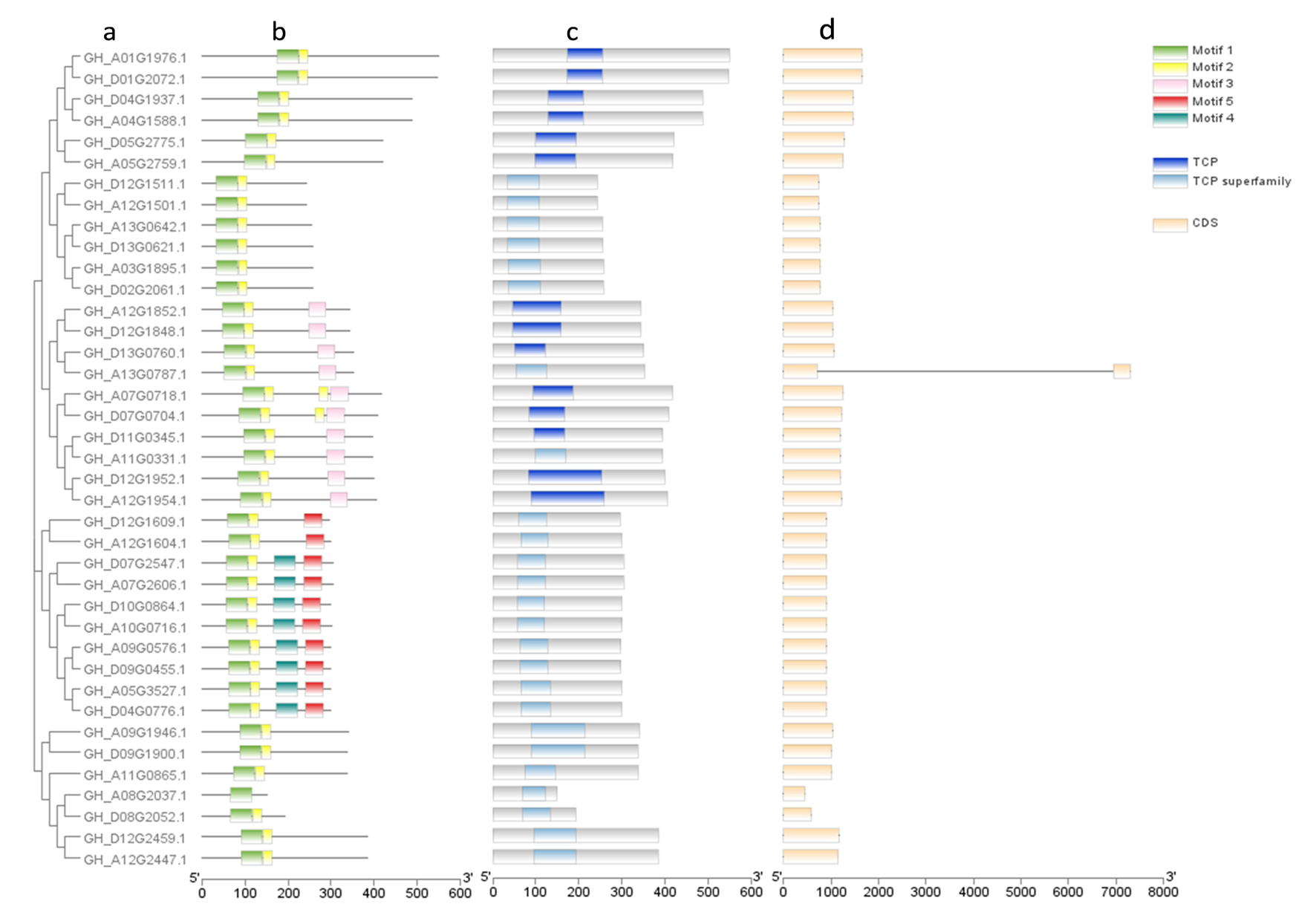
Fig.3 Phylogenetic tree, motif prediction, protein domain and gene structure of Class I TCP genes in upland cotton Note:a.Phylogenetic tree of TCP Class I genes; b.Motif prediction results of TCP Class I proteins; c.Protein domain of TCP Class I; d.Gene structure of Class I TCP
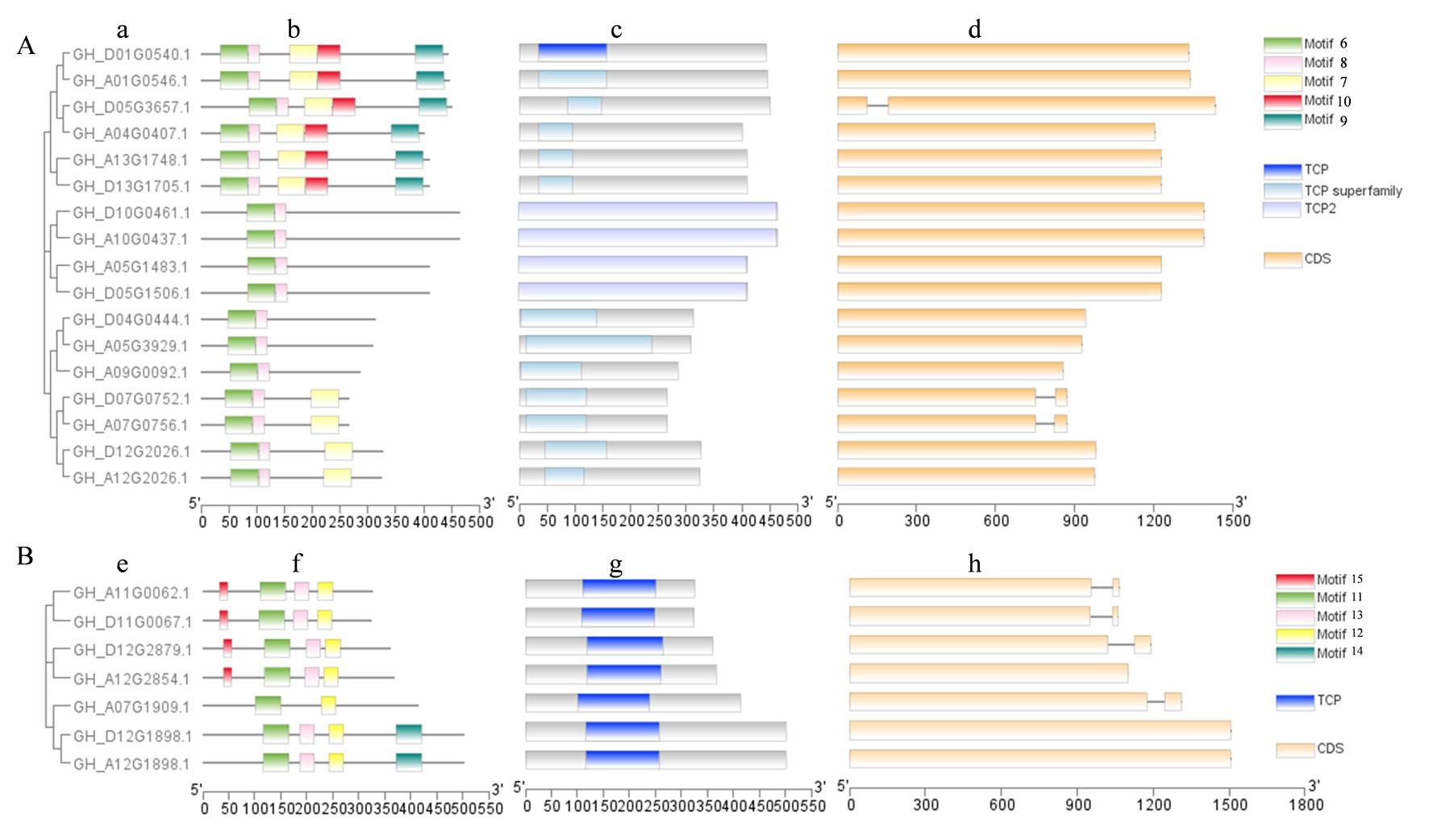
Fig.4 Phylogenetic tree, motif prediction, protein domain and gene structure of Class Ⅱ TCP genes in upland cotton Note:A.Phylogenetic tree, motif prediction, protein domain and gene structure of CIN genes in upland cotton; B.Phylogenetic tree, motif prediction, protein domain and gene structure of CYC/TB1 genes in upland cotton.a.Phylogenetic tree of CIN genes; b.Motif prediction results of CIN proteins; c.Protein domain of CIN; d.Gene structure of CIN; e.Phylogenetic tree of CYC/TB1 genes; f.Motif prediction results of CYC/TB1 proteins; g.Protein domain of CYC/TB1; h.Gene structure of CYC/TB1
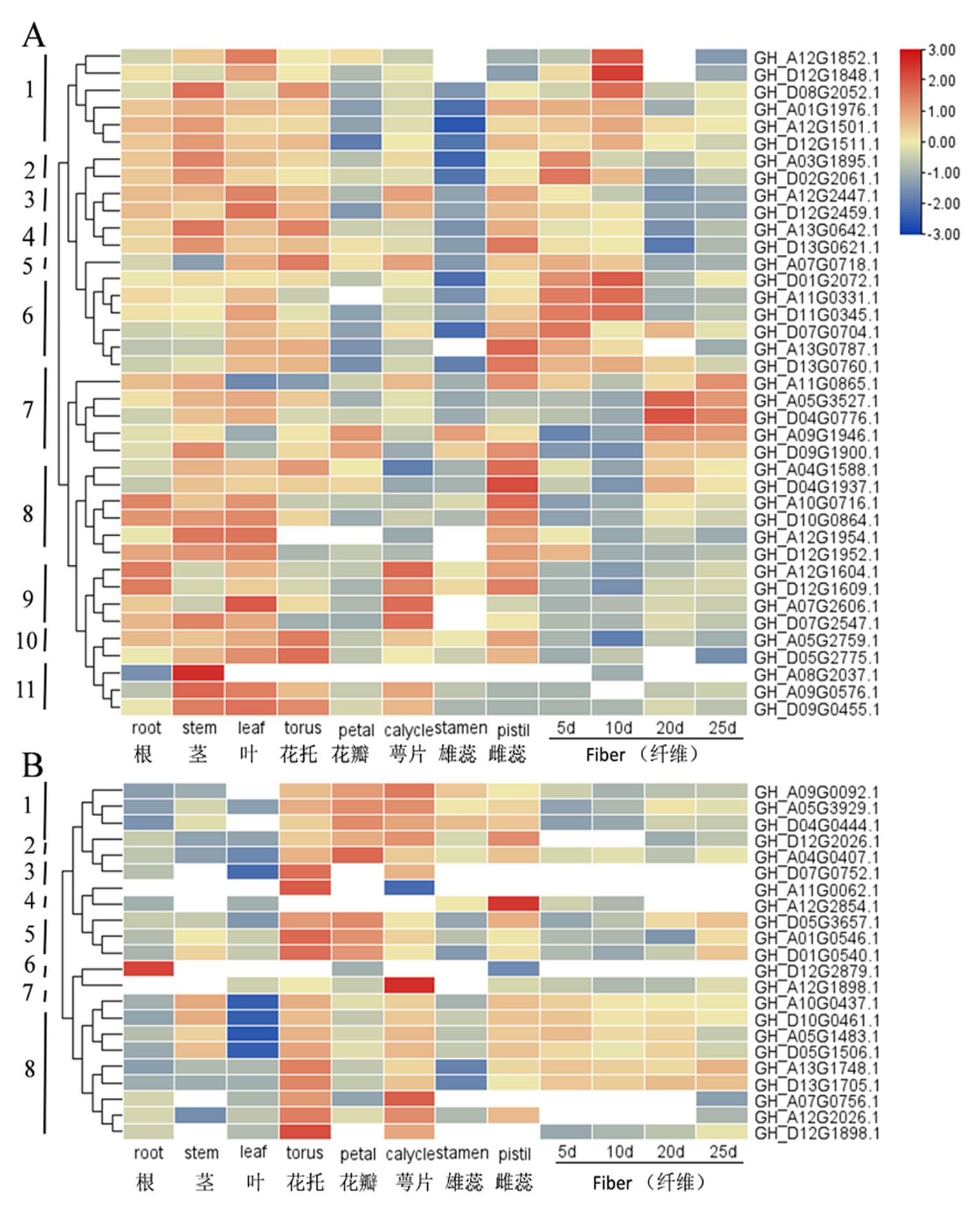
Fig.5 Expression analysis of TCP genes from different tissues in upland cotton Note:A: Expression analysis of Class I genes from different tissues in upland cotton; B: Expression analysis of Class Ⅱ genes from different tissues in upland cotton; The numerical range of the right scale represents the variation range of the expression level after normalization, with red as high expression level, blue as low expression level and white as missing value
| [1] |
Chai W, Jiang P, Huang G, et al. Identification and expression profiling analysis of TCP family genes involved in growth and development in maize[J]. Physiology and Molecular Biology of Plants, 2017, 23(4):779-791.
DOI URL |
| [2] |
Zhao J M, Zhai Z W, Li Y N, et al. Genome-wide identification and expression profiling of the TCP family genes in spike and grain development of wheat (Triticum aestivum L.)[J]. Front in Plant Science, 2018, 9:1282.
DOI URL |
| [3] | Cubas P. Role of TCP genes in the evolution of morphological characters in angiosperms. In: Cronk QCB, Bateman RM, Hawkins JA, eds. Developmental genetics and plant evolution[M]. London: Taylor & Francis, 2002, 65: 247-266. |
| [4] |
Riechmann J L, Heard J, Martin G, et al. Arabidopsis transcription factors: genome-wide comparative analysis among eukaryotes[J]. Science, 2000, 290(5499):2105-2110.
DOI PMID |
| [5] |
Ma J, Wang Q L, Sun R R, et al. Genome-wide identification and expression analysis of TCP transcription factors in Gossypium raimondii[J]. Scientific Reports, 2014, 4:6645.
DOI |
| [6] |
Navaud O, Dabos P, Carnus E, et al. TCP transcription factors predate the emergence of land plants[J]. Journal of Molecular Evolution, 2007, 65(1):23-33.
DOI URL |
| [7] |
Xiong Y Q, Liu T Y, Tian C G, et al. Transcription factors in rice: a genome-wide comparative analysis between monocots and eudicots[J]. Plant Molecular Biology, 2005, 59(1): 191-203.
DOI PMID |
| [8] |
Yao X, Ma H, Wang J, et al. Genome-wide comparative analysis and expression pattern of TCP gene families in Arabidopsis thaliana and Oryza sativa[J]. Journal of Integrative Plant Biology, 2007, 49(6): 885-897.
DOI URL |
| [9] | 梁欢欢. 陆地棉主要产量、品质性状鉴定与优异种质筛选[D]. 保定: 河北农业大学, 2018. |
| LIANG Huanhuan. Identification of main yield and quality traits of upland cotton and screening of elite germplasm resources[D]. Baoding: Hebei Agricultural University, 2018. | |
| [10] |
Cubas P, Lauter N, Doebley J, et al. The TCP domain: a motif found in proteins regulating plant growth and development[J]. The Plant Journal, 1999, 18: 215-222.
DOI URL |
| [11] |
Hao J, Tu L L, Hu H Y, et al. GbTCP, a cotton TCP transcription factor, confers fiber elongation and root hair development by a complex regulating system[J]. Journal of Experiment Botany, 2012, 63: 6267-6281.
DOI URL |
| [12] |
Li W, Li D D, Han L H, et al. Genome-wide identification and characterization of TCP transcription factor genes in upland cotton (Gossypium hirsutum)[J]. Scientific Reports, 2017, 7:10118.
DOI PMID |
| [13] | 杨菁玉. GhBIN2调控棉花株型发育机制的研究[D]. 郑州: 郑州大学, 2021. |
| YANG Jingyu. Regulation mechanism of GhBIN2 on plant type development in cotton[D]. Zhengzhou: Zhengzhou University, 2021. | |
| [14] | 刁阳阳. GhTIE1调节TCPs的转录活性控制棉花和拟南芥分枝发育[D]. 北京: 中国农业科学院, 2020. |
| DIAO Yangyang. GhTIE1 regulates branching through modulating the transcriptional activity of TCPs in cotton and Arabidopsis[D]. Beijing: Chinese Academy of Agricultural Sciences, 2020. | |
| [15] |
Wang M Y, Zhao P M, Cheng H Q, et al. The cotton transcription factor TCP14 functions in auxin-mediated epidermal cell differentiation and elongation[J]. Plant Physiology, 2013, 162: 1669-1680.
DOI URL |
| [16] |
Wang Y, Yu Y H, Chen Q J, et al. Heterologous expression of GbTCP4, a class II TCP transcription factor, regulates trichome formation and root hair development in Arabidopsis[J]. Genes, 2019, 10(9):726.
DOI URL |
| [17] |
Wang Y, Yu Y H, Wan H N, et al. The sea-island cotton GbTCP4 transcription factor positively regulates drought and salt stress responses[J]. Plant Science, 2022, 322:111329.
DOI URL |
| [18] | 韩利红. 棉花TCP转录因子的鉴定和特征分析及EIN3的功能研究[D]. 武汉: 华中师范大学, 2017. |
| HAN Lihong. Genome-wide identification and characterization of TCP transcription factors and study on the role of RIN3 in upland cotton (Gossypium hirsutum)[D]. Wuhan: Central China Normal University, 2017. | |
| [19] |
Hu Y, Chen JD, Fang L, et al. Gossypium barbadense and Gossypium hirsutum genomes provide insights into the origin and evolution of allotetraploid cotton[J]. Nature Genetics, 2019, 51(4): 739-748.
DOI |
| [20] | Ku C C. TCP6, a regulator in Arabidopsis gametophyte development and DNA damage response[D]. Edinburgh: University of Edinburgh, 2014. |
| [21] | Aguilar-Martínez JA, Sinha N. Analysis of the role of Arabidopsis class I TCP genes AtTCP7, AtTCP8, AtTCP22, and AtTCP23 in leaf development[J]. Frontiers in Plant Science, 2013, 4(406):1-13. |
| [22] |
Li X Y, Zhang G F, Liang Y H, et al. TCP7 interacts with Nuclear Factor-Ys to promote flowering by directly regulating SOC1 in Arabidopsis[J]. The Plant Journal, 2021, 108(5): 1493-1506.
DOI URL |
| [23] |
Danisman S, Wal F, Dhondt S, et al. Arabidopsis class I and class II TCP transcription factors regulate jasmonic acid metabolism and leaf development antagonistically[J]. Plant Physiology, 2012, 159:1511-1523.
DOI URL |
| [24] |
Viola I L, Manassero NGU, Ripoll R, et al. The Arabidopsis class I TCP transcription factor AtTCP11 is a developmental regulator with distinct DNA-binding properties due to the presence of a threonine residue at position 15 of the TCP domain[J]. Biochemical Journal, 2011, 435(1):143-155.
DOI URL |
| [25] |
Giraud E, Ng S, Carrie C, et al. TCP transcription factors link the regulation of genes encoding mitochondrial proteins with the circadian clock in Arabidopsis thaliana[J]. Plant Cell, 2010, 22: 3921-3934.
DOI URL |
| [26] | 安家兴. 拟南芥TCP11调控维管束的发育[D]. 兰州: 兰州大学, 2012. |
| AN Jiaxing. TCP11 regulates the development of vascular bundles in Arabidopsis thaliana[D]. Lanzhou: Lanzhou University, 2012. | |
| [27] |
Lopez JA, Sun Y L, Blair P B, et al. TCP three-way handshake: linking developmental processes with plant immunity[J]. Trends in Plant Science, 2015, 20(4):238-245.
DOI PMID |
| [28] |
Kieffer M, Master V, Waites R, et al. TCP14 and TCP15 affect internode length and leaf shape in Arabidopsis[J]. The Plant Journal, 2011, 68:147-158.
DOI PMID |
| [29] |
Davière J M, Wild M, Regnault T, et al. Class I TCP-DELLA interactions in inflorescence shoot apex determine plant height[J]. Current Biology, 2014, 24:1923-1928.
DOI URL |
| [30] |
Viola L, Camoirano A, Gonzalez D H. Redox-dependent modulation of anthocyanin biosynthesis by the TCP transcription factor TCP15 during exposure to high light intensity conditions in Arabidopsis[J]. Plant Physiology, 2016, 170:74-85.
DOI URL |
| [31] |
Uberti-Manassero N G, Lucero L E, Viola I L, et al. The class I protein AtTCP15 modulates plant development through a pathway that overlaps with the one affected by CIN-like TCP proteins[J]. Journal of Experiment Botany, 2012, 63: 809-823.
DOI URL |
| [32] | 张春雷. 拟南芥TCP15、TCP22基因功能的研究[D]. 兰州: 兰州大学. 2012. |
| ZHANG Chunlei. Functional analysis of two transcription factors, TCP15 and TCP22, from Arabidopsis thaliana[D]. Lanzhou: Lanzhou University, 2012. | |
| [33] |
Li ZY, Li B, Wu AD. The Arabidopsis transcription factor AtTCP15 regulates endoreduplication by modulating expression of key cell-cycle genes[J]. Molecular Plant, 2012, 5(1): 270-280.
DOI URL |
| [34] |
Takeda T, Amano K, Ohto M, et al. RNA interference of the Arabidopsis putative transcription factor TCP16 gene results in abortion of early pollen development[J]. Plant Molecular Biology, 2006, 61:165-177.
PMID |
| [35] |
Hervé C, Dabos P, Bardet C, et al. In vivo interference with AtTCP20 function induces severe plant growth alterations and deregulates the expression of many genes important for development[J]. Plant Physiology, 2009, 149: 1462-1477.
DOI URL |
| [36] |
Pruneda-Paz J L, Breton G, Para A, et al. A functional genomics approach reveals CHE as a component of the Arabidopsis circadian clock[J]. Science, 2009, 323(5920):1481-1485.
DOI PMID |
| [37] |
Cubas P, Coen E, Zapater JMM. Ancient asymmetries in the evolution of flowers[J]. Current Biology, 2001, 11(13):1050-1052.
PMID |
| [38] |
Guo Z X, Fujioka S, Miao S, et al. TCP1 modulates Brassinosteroid biosynthesis by regulating the expression of the key biosynthetic enzyme gene DWARF4 in Arabidopsis thaliana[J]. Plant Cell 2010, 22: 1161-1173.
DOI URL |
| [39] | An J X, Guo Z X, Gou X P, et al. TCP1 positively regulates the expression of DWF4 in Arabidopsis thaliana[J]. Plant Signaling & Behavior, 2011, 6: 1117-1118. |
| [40] |
Aguilar-Martínez J A, Poza-Carrión C, Cubas P. Arabidopsis BRANCHED1 acts as an integrator of branching signals within axillary buds[J]. Plant Cell, 2007, 19(2):458-472.
DOI PMID |
| [41] |
Palatnik J F, Allen E, Wu X L, et al. Control of leaf morphogenesis by microRNAs[J]. Nature, 2003, 425: 257-263.
DOI |
| [42] | 何志敏. 拟南芥转录因子TCP2与CRY1相互作用的遗传学与生化分析[D]. 长沙: 湖南大学, 2016. |
| HE Zhimin. The molecular genetical and biochemical study of transcription factor TCP2 and CRY1 interaction in Arabidopsis[D]. Changsha: Hunan University, 2016. | |
| [43] |
Koyama T, Furutani M, Tasaka M, et al. TCP transcription factors control the morphology of shoot lateral organs via negative regulation of the expression of boundary-specific genes in Arabidopsis[J]. Plant Cell, 2007, 19: 473-484.
DOI URL |
| [44] | Schommer C, Palatnik J F, Aggarwal P, et al. Control of jasmonate biosynthesis and senescence by miR319 targets[J]. PLoS Biology, 2008, 6(9): 1991-2001. |
| [45] |
Nag A, King S, Jack T. MiR319a targeting of TCP4 is critical for petal growth and development in Arabidopsis[J]. PNAS, 2009, 106(52):22534-22539.
DOI URL |
| [46] |
Zheng X H, Lan J Q, Yu H, et al. Arabidopsis transcription factor TCP4 represses chlorophyll biosynthesis to prevent petal greening[J]. Plant Communications, 2022, 3(4):100309.
DOI URL |
| [47] |
Zhou Y, Zhang D, An J, et al. TCP transcription factors regulate shade avoidance via directly mediating the expression of both PHYTOCHROME INTERACTING FACTORs and auxin biosynthetic genes.[J]. Plant Physiology, 2018, 176: 1850-1961.
DOI PMID |
| [48] |
Zhou Y, Xun Q Q, Zhang D Z, et al. TCP transcription factors associate with PHYTOCHROME INTERACTING FACTOR 4 and CRYPTOCHROME 1 to regulate thermomorphogenesis in Arabidopsis thaliana[J]. iScience, 2019, 15: 600-610.
DOI PMID |
| [49] | 周瑜. 拟南芥转录因子TCP17参与光信号转导和生长素合成[D]. 兰州: 兰州大学, 2013. |
| ZHOU Yu. Arabidopsis transcription factor TCP17 regulates light signaling transduction and auxin biosynthesis[D]. Lanzhou: Lanzhou University, 2013. | |
| [50] |
Zhu T, Liang CZ, Meng ZG, et al. CottonFGD: an integrated functional genomics database for cotton[J]. BMC Plant Biology, 2017, 17(1):101-109.
DOI PMID |
| [51] |
El-Gebali S, Mistry J, Bateman A, et al. The pfam protein families database in 2019[J]. Nucleic Acids Research, 2019, 47(Database issue): D427-D432.
DOI |
| [52] |
Finn R D, Clements J, Arndt W, et al. HMMER web server: 2015 update[J]. Nucleic Acids Research, 2015, 43(Web Server issue):W30-W38.
DOI URL |
| [53] |
Ivica L, Peer B. 20 years of the SMART protein domain annotation resource[J]. Nucleic Acids Research, 2018, 46(D1): D493-D496.
DOI URL |
| [54] | 张爱, 王彩香, 宿俊吉, 等. 陆地棉MADS-box家族基因鉴定及组织特异性表达分析[J]. 棉花学报, 2020, 32(5):404-417. |
| ZHANG Ai, WANG Caixia, SU Junjie, et al. Identification of MADS-box family and analysis of tissue specific expression in Gossypium hirsutum L.[J]. Cotton Sciences, 2020, 32(5):404-417. | |
| [55] |
Kumar S, Stecher G, Tamura K. MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets[J]. Molecular Biology and Evolution, 2016, 33(7): 1870-1874.
DOI PMID |
| [56] | Chen C J, Xia R, Chen H, et al. TBtools, a Tool kit for biologists integrating various HTS-data handling tools with a user-friendly interface[J]. bioRxiv, 2018, https://doi.org/10.1101/289660. |
| [57] | Bailey TL, Boden M, Buske FA, et al. MEME SUITE: tools for motif discovery and searching[J]. Nucleic Acids Research, 2009, 37(Web Server issue): W202-W208. |
| [58] |
Zhang T Z, Hu Y, Jiang W K, et al. Sequencing of allotetraploid cotton (Gossypium hirsutum L. acc. TM-1) provides a resource for fiber improvement[J]. Nature Biotechnology, 2015, 33:531-537.
DOI |
| [59] | 李燕, 姚金波, 陈伟, 等. 雷蒙德氏棉和亚洲棉TCP基因家族的生物信息学分析[J]. 棉花学报, 2016, 28(5):434-442. |
| LI Yan, YAO Jinbo, CHEN Wei, et al. Genome-wide identification of the TCP gene family in Gossypium raimondii and Gossypium arboreum using bioinformatic analysis[J]. Cotton Science, 2016, 28(5):434-442. |
| [1] | WANG Hui, GUO Jincheng, SONG Jia, ZHANG Tingjun, He Liangrong. Physiological and biochemical analysis of transgenic offspring of upland cotton GhCIPK6 under high temperature Stress [J]. Xinjiang Agricultural Sciences, 2023, 60(9): 2109-2119. |
| [2] | SANG Zhiwei, LIANG Yajun, GONG Zhaolong, ZHENG Juyun, WANG Junduo, LI Xueyuan, CHEN Quanjia. Analysis of mechanical harvesting characters of germplasm resources of different upland cotton [J]. Xinjiang Agricultural Sciences, 2023, 60(5): 1088-1098. |
| [3] | CHEN Liangliang, ZHANG Meng, GUO Liping, QI Tingxiang, ZHANG Xuexian, TANG Huini, WANG Hailin, QIAO Xiuqin, WU Jianyong, XING Chaozhu. Heterosis Performance and Their Parental Combining Ability Analysis of F1 and F2 Hybrids of Upland Cotton at Seedling Stage [J]. Xinjiang Agricultural Sciences, 2023, 60(2): 261-271. |
| [4] | LONG Tianyu, DENG Yahui, ZU Qianli, YANG Long, QU Yanying, CHEN Quanjia. Identification and Evaluation of Indoor Resistance to Cotton Fusarium wilt at Seedling Stage [J]. Xinjiang Agricultural Sciences, 2023, 60(2): 416-423. |
| [5] | LIU Chenxi, ZHU Yuting, ZHOU Qiang, CHEN Jin, ZHAO Wenjie, ZHENG Kai. Cloning and bioinformatics analysis of β-carotene isomerase GbD27-6 gene in sea island cotton [J]. Xinjiang Agricultural Sciences, 2023, 60(12): 2869-2877. |
| [6] | Mayila Yusuyin, YANG Ni, XU Haijiang, YANG Yanlong, ZHANG Dawei, LI Chunping, SHI Bixian, LAI Chengxia. Bioinformatics and expression analysis of UGT gene family of cotton glycosyltransferase damaged by Helicoverpa armigera [J]. Xinjiang Agricultural Sciences, 2023, 60(10): 2396-2403. |
| [7] | CAI Yi, DAI Dongyang, TAN Hai, WANG Di, YANG Fen, WANG Ling, SHENG Yunyan. Analysis of Genetic Structure and Construction of Gene Editing Vector of Melon AMS Gene [J]. Xinjiang Agricultural Sciences, 2023, 60(1): 61-68. |
| [8] | LI Min, MA Ying, Huercha , HE Wenwen, SHI Qianyun, Alimujiang Jiapaer, JIANG Qian, Bayinchahan . Expression of Dm86 Gene of Dermacenter Marginatus and Analysis of Immunogenicity [J]. Xinjiang Agricultural Sciences, 2022, 59(9): 2324-2332. |
| [9] | MA Xiaomei, LI Baocheng, DONG Chengguan, ZHOU Xiaofeng, WANG Xin, TIAN Qin, ZHAO Suqing, WANG Gang. Correlation between Leaf Morphological Indexes and Defoliation Regularity of Early-maturing Upland Cotton Varieties [J]. Xinjiang Agricultural Sciences, 2022, 59(6): 1301-1311. |
| [10] | MAO Tingyong, KONG Jie, HU Shoulin, ZHANG Wei, CHEN Jialin, LI Yanfang, WAN Sumei, CHEN Guodong. Morphological Comparison of Fiber Development in Different Upland Cotton Varieties in Southern Xinjiang [J]. Xinjiang Agricultural Sciences, 2022, 59(2): 279-290. |
| [11] | YANG Yanlong, MA Jun, SHI Weijun. Analysis of Genetic Diversity of Phenotypic Characters of Upland Cotton Germplasm Resources Imported [J]. Xinjiang Agricultural Sciences, 2022, 59(2): 310-319. |
| [12] | Rebiya Yusun, Wumaier Kurban, ZHANG Zhe, Maimaiti Moming, AI Xiantao. Analysis of Genetic Diversity of Main Characters in 288 Upland Cotton Germplasm Resources [J]. Xinjiang Agricultural Sciences, 2022, 59(12): 2879-2887. |
| [13] | DAI Qi, XU Ruiqiang, LI Ning, WANG Baike, WANG Juan, HUANG Shaoyong, Patiguli Aisimutuola, YU Qinghui. Genome-Wide Characterization and Analysis of GRF Gene Family in the Solanum pennellii [J]. Xinjiang Agricultural Sciences, 2022, 59(10): 2456-2465. |
| [14] | DENG Xiaojuan, SU Xiujuan, Mayila Yibulayin, LU Xiaoshuang, Maierdan Tuerxun, LIU Pengfei. Effects of Artificial Aging and Low-Temperature Stress on the Vigor of Upland Cotton Seed and Relevant Evaluation [J]. Xinjiang Agricultural Sciences, 2021, 58(6): 998-1005. |
| [15] | MA Xiaomei,LI Baocheng,WANG Xin,ZHAO Suqin,LIU Yongchang, HAN Huanyong, ZHOU Xiaofeng, DONG Chengguang. Interaction Effects of Early-maturing Upland Cotton Varieties and Meteorological Factors on Cotton Fiber Quality [J]. Xinjiang Agricultural Sciences, 2021, 58(2): 216-226. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||