

Xinjiang Agricultural Sciences ›› 2023, Vol. 60 ›› Issue (10): 2396-2403.DOI: 10.6048/j.issn.1001-4330.2023.10.007
• Crop Genetics and Breeding·Germplasm Resources·Molecular Genetics • Previous Articles Next Articles
Mayila Yusuyin( ), YANG Ni, XU Haijiang, YANG Yanlong, ZHANG Dawei, LI Chunping, SHI Bixian, LAI Chengxia(
), YANG Ni, XU Haijiang, YANG Yanlong, ZHANG Dawei, LI Chunping, SHI Bixian, LAI Chengxia( )
)
Received:2023-02-10
Online:2023-10-20
Published:2023-11-01
Correspondence author:
LAI Chengxia (1972-), female, native place: Urumqi, associate researcher, research direction: cotton molecular breeding, (E-mail)Supported by:
玛依拉·玉素音( ), 阳妮, 徐海江, 杨延龙, 张大伟, 李春平, 石必显, 赖成霞(
), 阳妮, 徐海江, 杨延龙, 张大伟, 李春平, 石必显, 赖成霞( )
)
通讯作者:
赖成霞(1972-),女,新疆乌鲁木齐人,副研究员,研究方向为棉花分子育种,(E-mail)作者简介:玛依拉·玉素音(1987-),女,新疆乌鲁木齐人,助理研究员,研究方向为棉花分子育种,(E-mail)119327132@qq.com
基金资助:CLC Number:
Mayila Yusuyin, YANG Ni, XU Haijiang, YANG Yanlong, ZHANG Dawei, LI Chunping, SHI Bixian, LAI Chengxia. Bioinformatics and expression analysis of UGT gene family of cotton glycosyltransferase damaged by Helicoverpa armigera[J]. Xinjiang Agricultural Sciences, 2023, 60(10): 2396-2403.
玛依拉·玉素音, 阳妮, 徐海江, 杨延龙, 张大伟, 李春平, 石必显, 赖成霞. 受棉铃虫诱导的棉花糖基转移酶UGT基因家族的生物信息学及表达分析[J]. 新疆农业科学, 2023, 60(10): 2396-2403.
Add to citation manager EndNote|Ris|BibTeX
URL: http://www.xjnykx.com/EN/10.6048/j.issn.1001-4330.2023.10.007
| 基因Gene | 正向引物Forward primer(5'-3') | 反向引物Reversr primer(5'-3') |
|---|---|---|
| histone 3 | GGTGGTGTGAAGAAGCCTCAT | AATTTCACGAACAAGCCTCTGGAA |
| UGTG01 | TTTCGCTCTGCAATAGGACA | AGCGATCTCCTTTAACTGACC |
| UGTG02 | ATCAAGGGACTAATGCGAAG | CAAATCCAAGCACCTAACGA |
| UGTGo4 | CTCAACCGTTTAACACCTCC | GCCTCTTCCTTTTACCCTT |
| UGTGo6 | ACGCTAAAATCCTTATTAACACC | AGTTAAAGAATGAAGATGGGACT |
| UGTGo7 | GTCATCTAAACTCTTTCCCACT | AGATTCAATAAAGGTCCCACT |
Tab.1 Primer sequences for RT-qPCR
| 基因Gene | 正向引物Forward primer(5'-3') | 反向引物Reversr primer(5'-3') |
|---|---|---|
| histone 3 | GGTGGTGTGAAGAAGCCTCAT | AATTTCACGAACAAGCCTCTGGAA |
| UGTG01 | TTTCGCTCTGCAATAGGACA | AGCGATCTCCTTTAACTGACC |
| UGTG02 | ATCAAGGGACTAATGCGAAG | CAAATCCAAGCACCTAACGA |
| UGTGo4 | CTCAACCGTTTAACACCTCC | GCCTCTTCCTTTTACCCTT |
| UGTGo6 | ACGCTAAAATCCTTATTAACACC | AGTTAAAGAATGAAGATGGGACT |
| UGTGo7 | GTCATCTAAACTCTTTCCCACT | AGATTCAATAAAGGTCCCACT |
| 基因 Gene | 蛋白长度 Length(aa) | 分子质量 Molecular weight (kDa) | 等电点 Theoretical pI | 不稳定系数 Instability index | 脂溶性指数 Aliphatic index | 亲水性 Hydrophilic |
|---|---|---|---|---|---|---|
| UGTG01 | 534 | 60 208.62 | 5.22 | 49.66 | 85.77 | -0.183 |
| UGTG02 | 469 | 52 972.26 | 5.01 | 48.29 | 86.84 | -0.202 |
| UGTGo4 | 493 | 54 852.32 | 5.58 | 40.18 | 93.75 | -0.143 |
| UGTGo6 | 469 | 52 660.09 | 5.31 | 40.46 | 84.58 | -0.233 |
| UGTGo7 | 468 | 52 743.1 | 4.97 | 49.85 | 89.12 | -0.166 |
Tab.2 Basic physical and chemical parameters of UGT protein of cotton glycosyltransferase
| 基因 Gene | 蛋白长度 Length(aa) | 分子质量 Molecular weight (kDa) | 等电点 Theoretical pI | 不稳定系数 Instability index | 脂溶性指数 Aliphatic index | 亲水性 Hydrophilic |
|---|---|---|---|---|---|---|
| UGTG01 | 534 | 60 208.62 | 5.22 | 49.66 | 85.77 | -0.183 |
| UGTG02 | 469 | 52 972.26 | 5.01 | 48.29 | 86.84 | -0.202 |
| UGTGo4 | 493 | 54 852.32 | 5.58 | 40.18 | 93.75 | -0.143 |
| UGTGo6 | 469 | 52 660.09 | 5.31 | 40.46 | 84.58 | -0.233 |
| UGTGo7 | 468 | 52 743.1 | 4.97 | 49.85 | 89.12 | -0.166 |
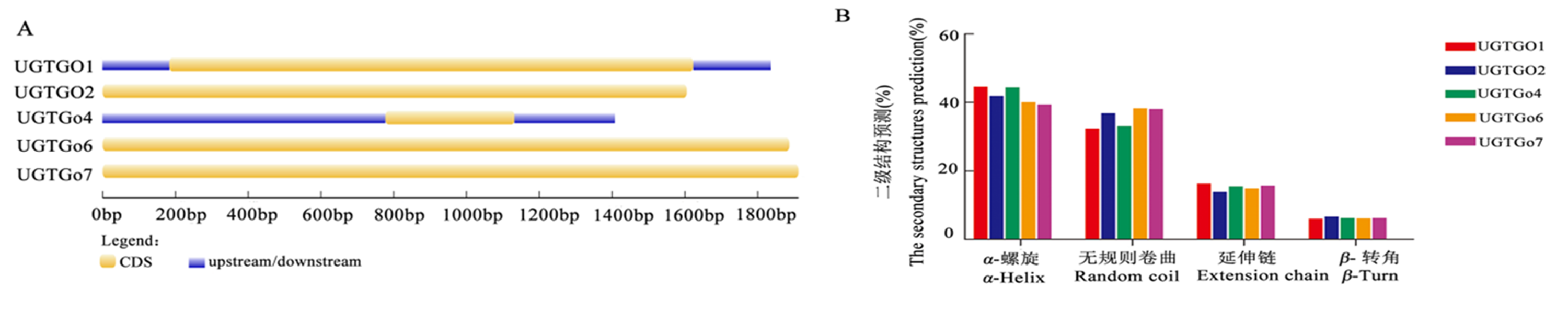
Fig.1 Gene structure analysis and the secondary structures prediction of cotton glycosyltransferase UGT family Note: A:the gene structure analysis of UGT gene family; B: the secondary structures prediction of UGT gene family
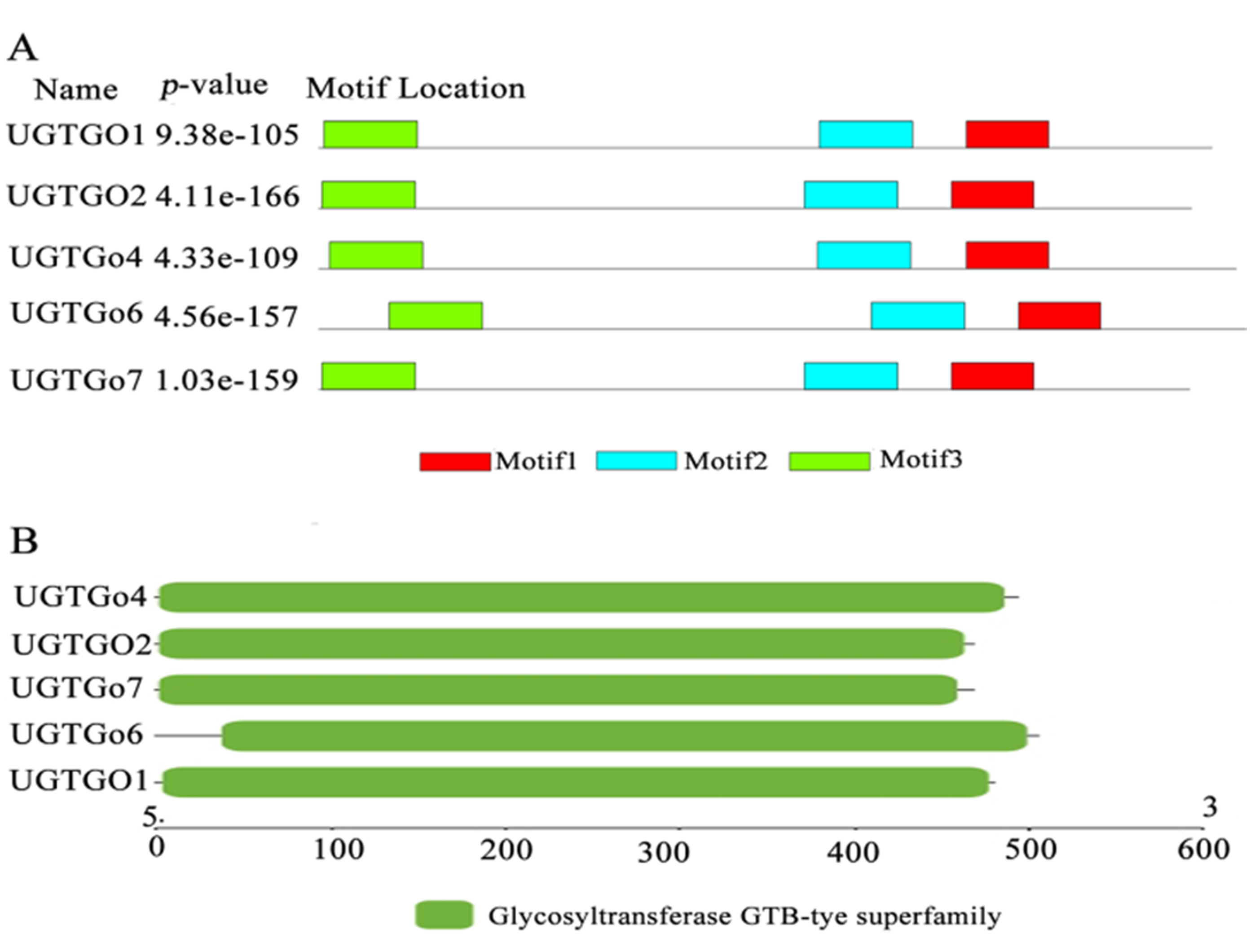
Fig.3 The analysis of protein conserved motifs and domain of cotton glycosyltransferase UGT gene family Note: A:protein conserved motif of UGT gene family;B.Conserved domain analysis of UGT gene family
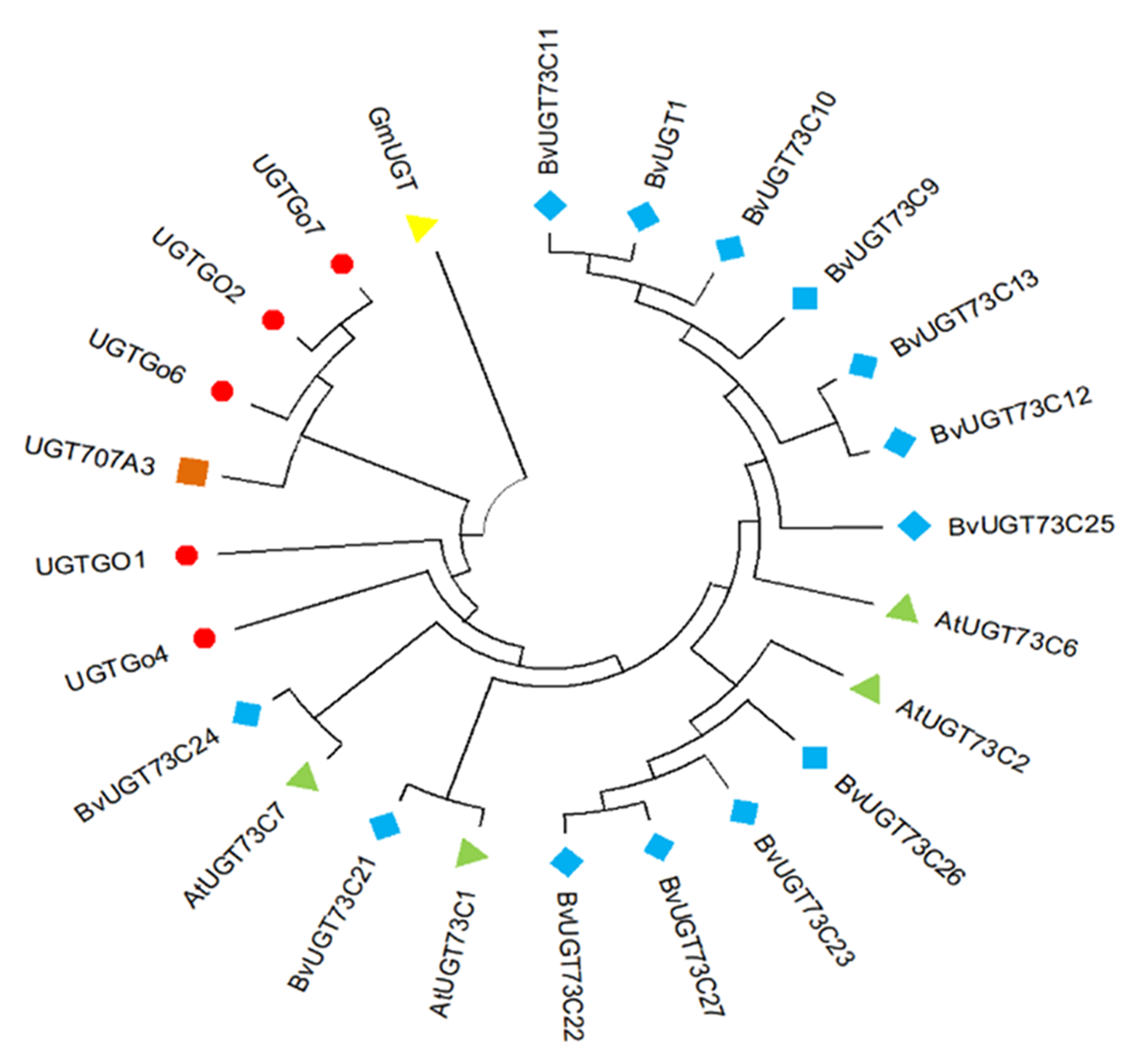
Fig.5 Phylogenetic tree of different plants UGTs Note:UGTs from the different species were labeled with different colored dots Arabidopsis thaliana, sorrel, rice, soybean, and cotton, represented as green triangles, blue diamonds, orange squares, yellow inverted triangles, and red circles, respectively.
| [1] |
Calvin W, Yang F, Brown SA, et al. Development of Economic Thresholds Toward Bollworm (Lepidoptera: Noctuidae), Management in Bt Cotton, and Assessment of the Benefits From Treating Bt Cotton With Insecticide[J]. Journal of Economic Entomology, 2021, 114(6): 2493-2504.
DOI PMID |
| [2] |
Chen B, Zhang Y, Sun Z, et al. Tissue-specific expression of GhnsLTPs identified via GWAS sophisticatedly coordinates disease and insect resistance by regulating metabolic flux redirection in cotton[J]. The Plant Journal, 2021, 107(3): 831-846.
DOI PMID |
| [3] | 于安东, 刘琳, 龙瑞才, 等. 植物UDP-糖基转移酶(UGT)的功能及应用前景[J]. 植物生理学报, 2022, 58(4): 631-642. |
|
YU Andong, LIU Lin, LONG Ruicai, et al. Function and application prospect of plant UDP-glycosyltransferase (UGT)[J]. Plant Physiology Journal, 2022, 58(4): 631-642.
DOI URL |
|
| [4] |
Rahimi S, Kim J, Mijakovic I, et al. Triterpenoid-biosynthetic UDP-glycosyltransferases from plants[J]. Biotechnology Advances, 2019, 37(7): 107394.
DOI URL |
| [5] |
Rehman H M, Nawaz M A, Shah Z H, et al. Comparative genomic and transcriptomic analyses of Family-1 UDP glycosyltransferase in three Brassica species and Arabidopsis indicates stress-responsive regulation[J]. Scientific reports, 2018, 8(1): 1875.
DOI PMID |
| [6] | Jing T, Zhang N, Gao T, et al. Glucosylation of (Z)-3-hexenol informs intraspecies interactions in plants: A case study in Camellia sinensis[J]. Plant, Cell & Environment. 2019, 42(4): 1352-1367. |
| [7] |
Augustin JM, Drok S, Shinoda T, et al. UDP-glycosyltransferases from the UGT73C subfamily in Barbarea vulgaris catalyze sapogenin 3-O-glucosylation in saponin-mediated insect resistance[J]. Plant Physiology, 160(4): 1881-1895.
DOI URL |
| [8] |
Zhang Y, Guo W, Chen L, et al. CRISPR/Cas9-Mediated Targeted Mutagenesis of GmUGT Enhanced Soybean Resistance Against Leaf-Chewing Insects Through Flavonoids Biosynthesis[J]. Frontiers in Plant Science, 2022, 13:802716.
DOI URL |
| [9] |
Yang Z, Li N, Kitano T, et al. Genetic mapping identifies a rice naringenin O-glucosyltransferase that influences insect resistance[J]. The Plant Journal, 2021, 106(5): 1401-1413.
DOI URL |
| [10] |
Gilbert MK, Bland JM, Shockey JM, et al. A Transcript Profiling Approach Reveals an Abscisic Acid-Specific Glycosyltransferase (UGT73C14) Induced in Developing Fiber of Ligon lintless-2 Mutant of Cotton (Gossypium hirsutum L.)[J]. PLoS ONE, 2013, 8(9): e75268.
DOI URL |
| [11] |
Qin L-X, Rao Y, Li L, et al. Cotton GalT1 Encoding a Putative Glycosyltransferase Is Involved in Regulation of Cell Wall Pectin Biosynthesis during Plant Development. PLoS ONE, 2013, 8(3): e59115.
DOI URL |
| [12] |
Krempl C, Sporer T, Reichelt M, et al. Potential detoxification of gossypol by UDP-glycosyltransferases in the two Heliothine moth species Helicoverpa armigera and Heliothis virescens[J]. Insect Biochemistry and Molecular Biology, 2016, 71:49-57.
DOI PMID |
| [13] |
Kosaka A, Manickavelu A, Kajihara D, et al. Altered gene expression profiles of wheat genotypes against Fusarium head blight[J]. Toxins (Basel), 2015, 7(2): 604-620.
DOI PMID |
| [14] |
Langlois-Meurinne M, Gachon CM, Saindrenan P. Pathogen-responsive expression of glycosyltransferase genes UGT73B3 and UGT73B5 is necessary for resistance to Pseudomonas syringae pv tomato in Arabidopsis[J]. Plant Physiology, 2005, 139(4): 1890-1901.
PMID |
| [15] |
Poppenberger B, Berthiller F, Lucyshyn D, et al. Detoxification of the Fusarium mycotoxin deoxynivalenol by a UDP-glucosyltransferase from Arabidopsis thaliana[J]. Journal of Biological Chemistry, 2003, 278(48): 47905-47914.
DOI PMID |
| [16] | 赖成霞, 玛依拉·玉素音, 高燕, 等. 藏红花酸糖基转移酶UGTCs4基因的克隆、生物信息学及表达分析[J]. 中草药, 2020, 51(4): 1044-1051. |
| LAI Chengxia, Mayila Yusuyin, GAO Yan, et al. Cloning, bioinformatics, expression mode analysis of crocetin glycosyltransferase UGTCs4[J]. Chinese Traditional and Herbal Drugs, 2020, 51(4): 1044-1051. | |
| [17] | 钟晓武, 付强, 邹颉, 等. 烟草糖基转移酶基因NtGT5a的克隆及原核表达. 烟草科技, 2014,(7): 69-74,84. |
| ZHONG Xiaowu, FU Qiang, ZOU Jie, et al. Cloning and Prokaryotic Expression of Glycosyltransferase Gene NtGT5a from Nicotiana tabacum[J]. Tobacco Science and Technology, 2014,(7): 69-74,84. | |
| [18] | 张建军, 文国琴, 刘震, 等. 拟南芥糖基转移酶基因UGT71C5启动子的克隆和功能分析. 四川大学学报(自然科学版), 2011, 48(4): 955-960. |
| ZHANG Jianjun, WEN Guoqin, LIU Zhen, et al. Cloning and functional analysis of UGT71C5 promoterin Arabidopsis thaliana[J]. Journal of Sichuan University (Natural Science Edition), 2011, 48(4): 955-960. | |
| [19] | 张永恩, 王群, 李潮海. 生物信息学在作物抗性研究中的应用展望[J]. 河南农业大学学报, 2005,(4): 25-28. |
| ZHANG Yongen, WANG Qun, LI Haichao. Outlook of the Applications of Bioinformatics in Crop Resistance Research[J]. Journal of Henan Agricultural University, 2005,(4): 25-28. | |
| [20] |
Huang Juan, Pang Chaoyou, Fan Shuli, et al. Genome-wide analysis of the family 1 glycosyltransferases in cotton.[J]. Molecular Genetics and Genomics : MGG, 2015, 290(5): 1805-1818.
DOI PMID |
| [21] | 翟琼, 郑浩霞, 胡博, 等. 花生DNA甲基转移酶基因家族生物信息学分析[J/OL]. 分子植物育种: 1-22[2022-11-19]. |
| ZHAI Qiong, ZHENG Haoxia, HU Bo, et al. Bioinformatics Analysis of Peanut DNA Methyltransferase Gene Family[J]. Molecular Plant Breeding, 1-22[2022-11-19]. | |
| [22] | 李寒, 夏鹏亮, 唐艳红, 等. 小麦OXR基因家族全基因组生物信息学及表达分析[J/OL]. 分子植物育种:1-10[2022-11-19]. |
| LI Han, XIA Pengliang, TANG Yanhong, et al. Genome-wide Bioinformatics and Expression Analysis of OXR Gene Family in Triticum Aestivum[J]. Molecular Plant Breeding, 1-10[2022-11-19]. | |
| [23] | 彭洪娴, 邱洁雅, 惠秋玲, 等. 资阳香橙砧木CjSPL基因家族鉴定及表达模式分析[J/OL]. 生物工程学报:1-19[2022-11-19]. |
| PENG Hongxian, IU Jieya, HUI Qiuling, et al. Identification of CjSPL gene family in Ziyang Xiangcheng rootstock and expression pattern analysis[J]. Chinese Journal of Biotechnology :1-19[2022-11-19]. | |
| [24] |
Wang D, Shi X, Liu D, et al. Transcriptome Profiling Revealed Potentially Critical Roles for Digestion and Defense-Related Genes in Insects' Use of Resistant Host Plants: A Case Study with Sitobion Avenae[J]. Insects, 2020, 11(2): 90.
DOI URL |
| [25] | 李健英. 棉花响应烟粉虱胁迫的分子机制及棉花体细胞胚胎表观基因组学研究[D]. 武汉: 华中农业大学, 2019. |
| LI Jianying. Molecular mechanism of cotton in response to whitefly ifestation and epigenomics basis of cotton somatic embryogenesis[D]. Wuhan: Huazhong Agricultural University, 2019. | |
| [26] | 陈云霄. 玉米和大刍草抗虫性评价及转录组分析[D]. 临汾: 山西师范大学, 2019. |
| CHEN Xiaoyun. Evaluation of Insect Resistance and Transcriptome Analysis of Maize and Teosinte[D]. Linfen: Journal of Shanxi Normal University, 2019. | |
| [27] | 刘龙博, 郑树轩, 郑洁. 石榴低温响应因子CBF基因家族鉴定及其表达分析[J]. 西北农业学报, 2022, 31(9): 1154-1167. |
| LIU Longbo, ZHENG Shuxuan, ZHENG Jie. Identification and Expression Analysis of CBF(C-repeat Binding Factor)Gene Family in Pomegranate(Punica granatum L.)[J]. Acta Agriculturae Boreali-Occidentalis Sinica, 2022, 31(9): 1154-1167. |
| [1] | LIU Chenxi, ZHU Yuting, ZHOU Qiang, CHEN Jin, ZHAO Wenjie, ZHENG Kai. Cloning and bioinformatics analysis of β-carotene isomerase GbD27-6 gene in sea island cotton [J]. Xinjiang Agricultural Sciences, 2023, 60(12): 2869-2877. |
| [2] | HU Wenran, ZHAO Zhun, SHAO Wukui, HUANG Quansheng. Identification of TCP family and analysis of tissue expression in upland cotton [J]. Xinjiang Agricultural Sciences, 2023, 60(11): 2627-2637. |
| [3] | CAI Yi, DAI Dongyang, TAN Hai, WANG Di, YANG Fen, WANG Ling, SHENG Yunyan. Analysis of Genetic Structure and Construction of Gene Editing Vector of Melon AMS Gene [J]. Xinjiang Agricultural Sciences, 2023, 60(1): 61-68. |
| [4] | LI Min, MA Ying, Huercha , HE Wenwen, SHI Qianyun, Alimujiang Jiapaer, JIANG Qian, Bayinchahan . Expression of Dm86 Gene of Dermacenter Marginatus and Analysis of Immunogenicity [J]. Xinjiang Agricultural Sciences, 2022, 59(9): 2324-2332. |
| [5] | DAI Qi, XU Ruiqiang, LI Ning, WANG Baike, WANG Juan, HUANG Shaoyong, Patiguli Aisimutuola, YU Qinghui. Genome-Wide Characterization and Analysis of GRF Gene Family in the Solanum pennellii [J]. Xinjiang Agricultural Sciences, 2022, 59(10): 2456-2465. |
| [6] | WANG Degang, MA Guanghuang, HE Pengpeng, XIONG Renci, CAO Yu, ZHANG Ping, HAN Xu, YANG Minglu. Gene Sequences and Expression Analysis of Fatty Acid Δ9 Desaturase of Euzophera pyriella during Different Stages [J]. Xinjiang Agricultural Sciences, 2020, 57(5): 877-887. |
| [7] | XU Hang, QUAN Shao-wen, MA Li, ZHOU Li, NIU Jian-xin. Molecular Cloning of the PsiERF Gene and Its Expression in Korla Fragrant Pears [J]. Xinjiang Agricultural Sciences, 2018, 55(9): 1616-1625. |
| [8] | HAO Xiao-yan, LI Jian-ping, CHANG Xiao-chun, Zumuremu, GAO Sheng-qi, CHEN Guo, HUANG Quan-sheng. Cloning and Bioinformatics Analysis of Salt-Tolerance of a PsaH Gene from Suaeda salsa [J]. Xinjiang Agricultural Sciences, 2018, 55(7): 1209-1217. |
| [9] | WANG Yu-kai, ZHAO Huan, NIU Jian-xin. Clonal Analysis of kfpSPL Gene and Its Promoter in Korla Fragrant Pear (Pyrus sinkiangensis) [J]. Xinjiang Agricultural Sciences, 2018, 55(6): 1017-1026. |
| [10] | HAO Xiao-yan, LI Jian-ping, Zumuremu Turxun, GAO Sheng-qi, HUANG Quan-sheng. Cloning and Bioinformatics Analysis of Salt-tolerance of PsaH Gene from Salicornia europaea [J]. Xinjiang Agricultural Sciences, 2017, 54(9): 1613-1620. |
| [11] | FENG Ya-ning;JIANG Zhi-qiang;MA Xiao-ling;LI Jiang-wei. Analysis of the Diversity of VHH Library from Bactrian Camel (Camelus bactrianus) with Bioinformatics [J]. , 2016, 53(8): 1533-1538. |
| [12] | ZHANG Jin-bo;LI Bo;YANG Yang;HU Wen-ran;CHEN Fan-yuan;XIE Li-xia;FAN Ling. Genomi-wide Investigation and Expression Analysis of CAD Gene Family in Gossypium hirsutum [J]. , 2016, 53(7): 1177-1187. |
| [13] | GUO Chun-miao;YANG Bo;Tudi Maimaiti;LI Ning;TANG Ya-ping;GONG Peng;WANG Ji-xun. Studies on Polymorphism and Bioinformatics of 8 Xinjiang Almonds Self-incompatibility Gene (S-gene) [J]. , 2016, 53(6): 1067-1073. |
| [14] | . Bioinformatics Analysis of PSY in 21 Plant Species [J]. , 2015, 52(12): 2157-2165. |
| [15] | SU Li-bo;Ailizati Aili;LIU Liu;PENG Dan;YANG Yi;ZHANG Fu-chun. Cloning and Expression Analysis of Salt-stress Responsive Hc2a1 Gene from Halostachys caspica [J]. , 2014, 51(9): 1686-1691. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||