

Xinjiang Agricultural Sciences ›› 2022, Vol. 59 ›› Issue (10): 2456-2465.DOI: 10.6048/j.issn.1001-4330.2022.10.014
• Horticultural Special Local Products·Forestry·Agricultural Product Processing Engineering • Previous Articles Next Articles
DAI Qi1( ), XU Ruiqiang1, LI Ning1,2, WANG Baike1, WANG Juan1, HUANG Shaoyong2, Patiguli Aisimutuola1, YU Qinghui1(
), XU Ruiqiang1, LI Ning1,2, WANG Baike1, WANG Juan1, HUANG Shaoyong2, Patiguli Aisimutuola1, YU Qinghui1( )
)
Received:2021-11-30
Online:2022-10-20
Published:2022-12-21
Correspondence author:
YU Qinghui
Supported by:
戴麒1( ), 徐瑞强1, 李宁1,2, 王柏柯1, 王娟1, 黄少勇2, 帕提古丽·艾斯木托拉1, 余庆辉1(
), 徐瑞强1, 李宁1,2, 王柏柯1, 王娟1, 黄少勇2, 帕提古丽·艾斯木托拉1, 余庆辉1( )
)
通讯作者:
余庆辉
作者简介:戴麒(1984-)男,辽宁人,助理研究员,研究方向为番茄遗传育种,(E-mail) daiqixj@163.com
基金资助:CLC Number:
DAI Qi, XU Ruiqiang, LI Ning, WANG Baike, WANG Juan, HUANG Shaoyong, Patiguli Aisimutuola, YU Qinghui. Genome-Wide Characterization and Analysis of GRF Gene Family in the Solanum pennellii[J]. Xinjiang Agricultural Sciences, 2022, 59(10): 2456-2465.
戴麒, 徐瑞强, 李宁, 王柏柯, 王娟, 黄少勇, 帕提古丽·艾斯木托拉, 余庆辉. 潘那利番茄GRF基因家族全基因组鉴定分析[J]. 新疆农业科学, 2022, 59(10): 2456-2465.
Add to citation manager EndNote|Ris|BibTeX
URL: http://www.xjnykx.com/EN/10.6048/j.issn.1001-4330.2022.10.014
| 基因名 name | 基因 id | 染色体 Chromosome | 氨基酸数目 Number of amino acids (aa) | 分子量 MW (Da) | 等电点 pI | 亲水性 (GRAVY) | 脂肪 指数 Aliphatic index | 亚细胞定位 Subcellular localization |
|---|---|---|---|---|---|---|---|---|
| SpGRF1 | Sopen01g037310 | Spenn-ch01 | 420 | 46 630.8 | 9.31 | -0.755 | 61.26 | N,Cy,Ch,E |
| SpGRF2 | Sopen02g036640 | Spenn-ch02 | 604 | 64 906.7 | 8.23 | -0.504 | 62.4 | N,Cy,Ch |
| SpGRF3 | Sopen03g022150 | Spenn-ch03 | 461 | 50 686.8 | 6.69 | -0.582 | 60.74 | N,Ch |
| SpGRF4 | Sopen04g031110 | Spenn-ch04 | 595 | 64 143.4 | 8.22 | -0.649 | 58.27 | N,Cy,PM |
| SpGRF5 | Sopen07g021120 | Spenn-ch07 | 339 | 38 958.1 | 9.07 | -0.932 | 47.46 | N |
| SpGRF6 | Sopen08g001470 | Spenn-ch08 | 356 | 39 157 | 8.73 | -0.725 | 48.71 | N |
| SpGRF7 | Sopen08g024600 | Spenn-ch08 | 361 | 39 825.8 | 8.13 | -0.753 | 50.25 | N |
| SpGRF8 | Sopen08g028160 | Spenn-ch08 | 202 | 23 232.7 | 9.31 | -0.621 | 66.53 | N |
| SpGRF9 | Sopen08g031420 | Spenn-ch08 | 385 | 42 692.3 | 8.48 | -0.788 | 53.25 | N |
| SpGRF10 | Sopen10g032980 | Spenn-ch10 | 388 | 42 247.5 | 9.27 | -0.659 | 57.86 | N |
Table 1 SpGRFs gene family physicochemical properties
| 基因名 name | 基因 id | 染色体 Chromosome | 氨基酸数目 Number of amino acids (aa) | 分子量 MW (Da) | 等电点 pI | 亲水性 (GRAVY) | 脂肪 指数 Aliphatic index | 亚细胞定位 Subcellular localization |
|---|---|---|---|---|---|---|---|---|
| SpGRF1 | Sopen01g037310 | Spenn-ch01 | 420 | 46 630.8 | 9.31 | -0.755 | 61.26 | N,Cy,Ch,E |
| SpGRF2 | Sopen02g036640 | Spenn-ch02 | 604 | 64 906.7 | 8.23 | -0.504 | 62.4 | N,Cy,Ch |
| SpGRF3 | Sopen03g022150 | Spenn-ch03 | 461 | 50 686.8 | 6.69 | -0.582 | 60.74 | N,Ch |
| SpGRF4 | Sopen04g031110 | Spenn-ch04 | 595 | 64 143.4 | 8.22 | -0.649 | 58.27 | N,Cy,PM |
| SpGRF5 | Sopen07g021120 | Spenn-ch07 | 339 | 38 958.1 | 9.07 | -0.932 | 47.46 | N |
| SpGRF6 | Sopen08g001470 | Spenn-ch08 | 356 | 39 157 | 8.73 | -0.725 | 48.71 | N |
| SpGRF7 | Sopen08g024600 | Spenn-ch08 | 361 | 39 825.8 | 8.13 | -0.753 | 50.25 | N |
| SpGRF8 | Sopen08g028160 | Spenn-ch08 | 202 | 23 232.7 | 9.31 | -0.621 | 66.53 | N |
| SpGRF9 | Sopen08g031420 | Spenn-ch08 | 385 | 42 692.3 | 8.48 | -0.788 | 53.25 | N |
| SpGRF10 | Sopen10g032980 | Spenn-ch10 | 388 | 42 247.5 | 9.27 | -0.659 | 57.86 | N |

Fig.4 SpGRFs gene family conserved motifs and structural analysis Note:A.Distribution of the conserved motifs.B.Exon/intron distribution of SpGRF genes.The same colored boxes represent the same motifs
| 重复基因对 Duplicated Gene Pairs | 同义替换率 Ks | 非同义替换率Ka | Ka/Ks | 复制类型 Duplicated Type | 分化时间 Time(Mya*) |
|---|---|---|---|---|---|
| SpGRF6/SpGRF7 | 1.077 | 0.979 | 0.909 | Segmental | 2.835 |
| SpGRF2/SpGRF4 | 1.006 | 0.998 | 0.992 | Segmental | 2.648 |
| SpGRF3/SpGRF9 | 0.869 | 1.035 | 1.191 | Segmental | 2.287 |
Table 2 Analysis of spgrf paralog KaKs and calculation of differentiation time
| 重复基因对 Duplicated Gene Pairs | 同义替换率 Ks | 非同义替换率Ka | Ka/Ks | 复制类型 Duplicated Type | 分化时间 Time(Mya*) |
|---|---|---|---|---|---|
| SpGRF6/SpGRF7 | 1.077 | 0.979 | 0.909 | Segmental | 2.835 |
| SpGRF2/SpGRF4 | 1.006 | 0.998 | 0.992 | Segmental | 2.648 |
| SpGRF3/SpGRF9 | 0.869 | 1.035 | 1.191 | Segmental | 2.287 |
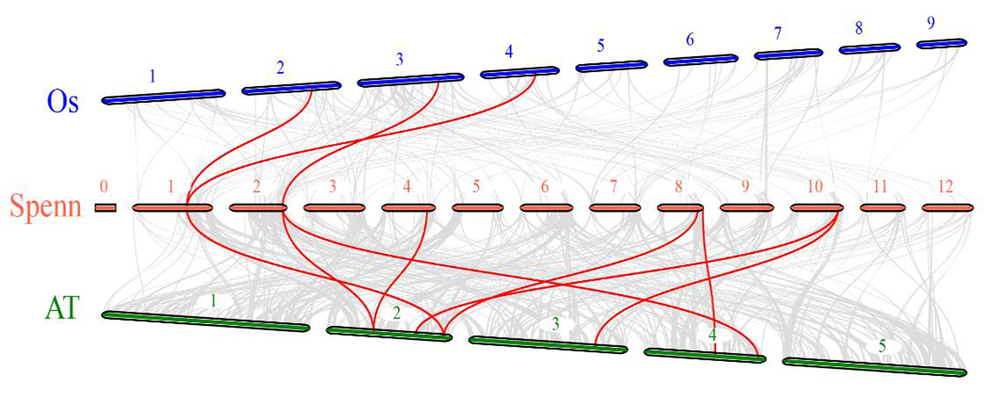
Fig.8 SpGRFs gene families Interspecies colinearity analysis Note:Genomes of rice(blue),panari tomato(red) and Arabidopsis(green) from top to bottom,The red line connects the collinearity gene
| [1] |
Mohammad A O, Sebastian P, Ushio F, et al. Growth-Regulating Factors(GRFs)a small transcription factor family with important functions in plant biology[J]. Molecular Plant, 2015, 8(7):998-1010.
DOI URL |
| [2] |
Zhang D F, Bo L, Jia G Q, et al. Isolation and characterization of genes encoding GRF transcription factors and GIF transcriptional coactivators in Maize(Zea mays L.)[J]. Plant Science, 2008, 175(6):809-817.
DOI URL |
| [3] |
Büyük I, Aras S. Genome-wide in silico identification characterization and transcriptional analysis of the family of growth-regulating factors in common bean(Phaseolus vulgaris L.)subjected to polyethylene glycol-induced drought stress[J]. Archives of Biological Sciences, 2017, 69(1):5-14
DOI URL |
| [4] | Kim J H, Kende H. A transcriptional coactivator AtGIF1 is involved in regulating leaf growth and morphology in Arabidopsis[J]. PANS, 2004, 101(36):13374-13379 |
| [5] |
Abdul F A, Muhammad S, Nazaruddin N, et al. MicroRNA and Transcription Factor:Key Players in Plant Regulatory Network[J]. Frontiers in Plant Science, 2017, 8:565.
DOI PMID |
| [6] |
Choi D, Kim J H, Kende H, et al. Whole genome analysis of the OsGRF gene family encoding plant-specific putative transcription activators in rice(Oryza sativa L.)[J]. Plant and Cell Physiology, 2004, 45(7):897-904.
DOI URL |
| [7] |
Khatun K, Robin A H K, Park J I, et al. Molecular characterization and expression profiling of tomato GRF transcription factor family genes in response to abiotic stresses and phytohormones[J]. International Journal of Molecular Science, 2017, 18(5):1056.
DOI URL |
| [8] | 周蕾. 烟草叶片发育相关ARF和GRF家族基因的表达分析与功能验证[D]. 北京: 中国农业科学院博士论文, 2020. |
| ZHOU Lei. Expression analysis and functional verification of ARF and GRF family genes related to tobacco leaf development[D]. Beijing: Chinese Academy of Agricultural Sciences, 2020. | |
| [9] |
Knaap E, Kim J H, et al. A novel gibberellin-induced gene from rice and its potential regulatory role in stem growth[J]. Plant Physiology, 2000, 122(3):695-704.
PMID |
| [10] |
Lee B H, Ko J H, Lee S, et al. The Arabidopsis GRF-INTERACTING FACTOR gene family performs an overlapping function in determining organ size as well as multiple developmental properties[J]. Plant Physiology, 2009, 151(2):655-668.
DOI URL |
| [11] | Cao Y, Han Y, Jin Q, et al. Comparative genomic analysis of the GRF genes in Chinese pear(Pyrus bretschneideri Rehd),poplar(Populous),grape(Vitis vinifera),Arabidopsis and rice(Oryza sativa)[J]. Frontiers in Plant Science, 2016, 7:1750. |
| [12] |
Cao J F, Huang J Q, Liu X, et al. Genome-wide characterization of the GRF family and their roles in response to salt stress in Gossypium[J]. BMC genomics, 2020, 21(1):1-16.
DOI URL |
| [13] | 王晓芳, 昌秦湘, 王淑芳, 等. 葡萄GRF基因家族的生物信息学分析[J]. 分子植物育种, 2021, 19(18):5975-5983. |
| WANG Xiaofang, CHANG Qinxiang, WANG Shufang, et al. Bioinformatics analysis of grape GRF gene family[J]. Molecular plant breeding, 2021, 19(18):5975-5983. | |
| [14] |
Ma J Q, Jian H J, Yang B, et al. Genome-wide analysis and expression profiling of the GRF gene family in oilseed rape(Brassica napus L.)[J]. Gene, 2017, 620:36-45.
DOI URL |
| [15] |
Kim J H, Lee B H. Growth-regulating FACTOR4 of Arabidopsis thaliana is required for development of leaves,cotyledons,and shoot apical meristem[J]. Journal of Plant Biology, 2006, 49(6):463-468.
DOI URL |
| [16] |
Kim J H, Tsukaya H. Regulation of plant growth and development by the GROWTH-REGULATING FACTOR and GRF-INTERACTING FACTOR duo[J]. Journal of Experimental Botany, 2015, 66(20):6093-6107.
DOI PMID |
| [17] |
Hewezi T, Maier T R, Nettleton D, et al. The Arabidopsis microRNA396-GRF1/GRF3 regulatory module acts as a developmental regulator in the reprogramming of root cells during cyst nematode infection[J]. Plant physiology, 2012, 159(1):321-335.
DOI PMID |
| [18] |
Kim J S, Mizoi J, Kidokoro S, et al. Arabidopsis GROWTH-REGULATING FACTOR7 functions as a transcriptional repressor of abscisic acid and osmotic stress-responsive genes,including DREB2A[J]. The Plant Cell, 2012, 24(8):3393-3405.
DOI URL |
| [19] |
Zhou Y, Ge L, Li G, et al. Characterization and expression analysis of growth regulating factor(GRF)family genes in cucumber[J]. Archives of Biological Sciences, 2018, 70(4):629-637.
DOI URL |
| [20] |
Wang F, Qiu N, Ding Q, et al. Genome-wide identification and analysis of the growth-regulating factor family in Chinese cabbage(Brassica rapa L.)[J]. BMC Genomics, 2014, 15(1):1-12.
DOI URL |
| [21] | 林涛. 番茄基因组多样性和演化的遗传学基础[D]. 北京: 中国农业大学博士论文, 2016. |
| LIN Tao. Genetic Basis of Tomato Genome Diversity and Evolution[D]. Beijing: China Agricultural University, 2016. | |
| [22] |
Khatun K, Robin A H K, Park J I, et al. Molecular characterization and expression profiling of tomato GRF transcription factor family genes in response to abiotic stresses and phytohormones[J]. International journal of molecular sciences, 2017, 18(05):1056.
DOI URL |
| [23] | 袁岐, 张春利, 赵婷婷, 等. 番茄GRF转录因子家族的生物信息学分析[J]. 分子植物育种, 2017, 15(8):2949-2956. |
| YUAN Qi, ZHANG Chunli, ZHAO Tingting, et al. Bioinformatics analysis of tomato GRF transcription factor family[J]. Molecular Plant Breeding, 2017, 15(8):2949-2956. | |
| [24] | 娄永峰, 宋晓琛, 孙化雨, 等. 毛竹PeGRF1基因克隆及初步功能分析[J]. 分子植物育种, 2021, 19(21):7084-7091. |
| LOU Yongfeng, SONG Xiaochen, SUN Huayu, et al. Cloning and preliminary functional analysis of PeGRF1 gene from Phyllostachys edulis[J]. Molecular Plant Breeding, 2021, 19(21):7084-7091. | |
| [25] | Bailey T L, Boden M, Buske F A, et al. MEME SUITE:tools for motif discovery and searching[J]. Nucleic acids research, 2009, 37(2):202-208. |
| [26] |
Guindon S, Gascuel O. A simple fast and accurate algorithm to estimate large phylogenies by maximum likelihood[J]. Systematic biology, 2003, 52(5):696-704.
DOI URL |
| [27] | 于继高. 葫芦科作物基因组形成过程的比较分析[D]. 唐山: 华北理工大学硕士论文, 2020. |
| YU Jigao. Genetic Basis of Tomato Genome Diversity and Evolution[D]. Tangshan: North China University of Science and Technology, 2020. | |
| [28] | 刘坤, 齐静, 刘梦兰, 等. 芝麻GRF基因家族全基因组鉴定分析[J]. 分子植物育种, 2020, 18(22):7323-7333. |
| LIU Kun, QI Jing, LIU Menglan, et al. Genome-wide identification of sesame GRF gene family[J]. Molecular Plant Breeding, 2020, 18(22):7323-7333. | |
| [29] | 赵慧霞, 李凤霞, 郭瑞, 等. 辣椒属GRF基因家族全基因组鉴定和表达分析[J]. 园艺学报, 2020, 47(11):2145-2160. |
| ZHAO Huixia, LI Fengxia, GUO Rui, et al. Genome-wide Identification and Expression Analysis of GRF Gene Family in Capsicum[J]. Acta Horticulturae Sinica, 2020, 47(11):2145-2160. | |
| [30] | 阮城城, 胡福初, 王祥和, 等. 基于转录组的荔枝GRF基因家族的鉴定及表达分析[J]. 分子植物育种, 2021, 19(20):6671-6681. |
| RUAN Chengcheng, HU Fuchu, WANG Xianghe, et al. Identification and Expression Analysis of GRF Gene Family in Litchi Based on Transcriptome[J]. Molecular Plant Breeding, 2021, 19(20):6671-6681. | |
| [31] | 张志康, 郭绩宁, 郭之乐, 等. 芥菜GRF转录因子家族的全基因组鉴定与分析[J]. 分子植物育种, 2020, 18(15):4878-4885. |
| ZHANG Zhikang, Guo Jining, GUO Zhile, et al. Genome-wide identification and analysis of mustard GRF transcription factor family[J]. Molecular Plant Breeding, 2020, 18(15):4878-4885. | |
| [32] |
Kim J H, Choi D, Kende H. The AtGRF family of putative transcription factors is involved in leaf and cotyledon growth in Arabidopsis[J]. The Plant Journal, 2003, 36(1):94-104.
DOI URL |
| [33] | 田娜, 刘范, 伍俊为, 等. 香蕉GRF基因家族的全基因组鉴定及表达分析[J]. 果树学报, 2020, 37(12):1821-1835. |
| TIAN Na, LIU Fan, WU Junwei, et al. Genome-wide identification and expression analysis of banana GRF gene family[J]. Journal of Fruit Science, 2020, 37(12):1821-1835. | |
| [34] |
马超, 原佳乐, 张苏, 等. GRF转录因子对植物生长发育及胁迫响应调控的分子机制[J]. 核农学报, 2017, 31(11):2145-2153.
DOI |
| MA Chao, YUAN Jiale, ZHANG Su, et al. Molecular Mechanism of GRF Transcription Factor Regulating Plant Growth and Stress Response[J]. Acta Agriculturae Nucleare Sinica, 2017, 31(11):2145-2153. | |
| [35] |
Chen F, Yang Y, Luo X, et al. Genome-wide identification of GRF transcription factors in soybean and expression analysis of GmGRF family under shade stress[J]. BMC plant biology, 2019, 19(1):1-13.
DOI URL |
| [36] | 方荣, 陈学军, 缪南生, 等. 茄科植物比较基因组学研究进展[J]. 江西农业学报, 2007,(2):35-38,45. |
| FANG Rong, CHEN Xujun, MIAO Nanshen, et al. Advances in Comparative Genomics of Solanaceae[J]. Acta Agriculture Jiangxi, 2007,(2):35-38,45. | |
| [37] |
Rodriguez R E, Mecchia M A, Debernardi J M, et al. Control of cell proliferation in Arabidopsis thaliana by microRNA miR396[J]. Development, 2010, 137(1):103-112.
DOI PMID |
| [1] | LIU Chenxi, ZHU Yuting, ZHOU Qiang, CHEN Jin, ZHAO Wenjie, ZHENG Kai. Cloning and bioinformatics analysis of β-carotene isomerase GbD27-6 gene in sea island cotton [J]. Xinjiang Agricultural Sciences, 2023, 60(12): 2869-2877. |
| [2] | HU Wenran, ZHAO Zhun, SHAO Wukui, HUANG Quansheng. Identification of TCP family and analysis of tissue expression in upland cotton [J]. Xinjiang Agricultural Sciences, 2023, 60(11): 2627-2637. |
| [3] | Mayila Yusuyin, YANG Ni, XU Haijiang, YANG Yanlong, ZHANG Dawei, LI Chunping, SHI Bixian, LAI Chengxia. Bioinformatics and expression analysis of UGT gene family of cotton glycosyltransferase damaged by Helicoverpa armigera [J]. Xinjiang Agricultural Sciences, 2023, 60(10): 2396-2403. |
| [4] | CAI Yi, DAI Dongyang, TAN Hai, WANG Di, YANG Fen, WANG Ling, SHENG Yunyan. Analysis of Genetic Structure and Construction of Gene Editing Vector of Melon AMS Gene [J]. Xinjiang Agricultural Sciences, 2023, 60(1): 61-68. |
| [5] | LI Min, MA Ying, Huercha , HE Wenwen, SHI Qianyun, Alimujiang Jiapaer, JIANG Qian, Bayinchahan . Expression of Dm86 Gene of Dermacenter Marginatus and Analysis of Immunogenicity [J]. Xinjiang Agricultural Sciences, 2022, 59(9): 2324-2332. |
| [6] | WANG Degang, MA Guanghuang, HE Pengpeng, XIONG Renci, CAO Yu, ZHANG Ping, HAN Xu, YANG Minglu. Gene Sequences and Expression Analysis of Fatty Acid Δ9 Desaturase of Euzophera pyriella during Different Stages [J]. Xinjiang Agricultural Sciences, 2020, 57(5): 877-887. |
| [7] | XU Hang, QUAN Shao-wen, MA Li, ZHOU Li, NIU Jian-xin. Molecular Cloning of the PsiERF Gene and Its Expression in Korla Fragrant Pears [J]. Xinjiang Agricultural Sciences, 2018, 55(9): 1616-1625. |
| [8] | HAO Xiao-yan, LI Jian-ping, CHANG Xiao-chun, Zumuremu, GAO Sheng-qi, CHEN Guo, HUANG Quan-sheng. Cloning and Bioinformatics Analysis of Salt-Tolerance of a PsaH Gene from Suaeda salsa [J]. Xinjiang Agricultural Sciences, 2018, 55(7): 1209-1217. |
| [9] | WANG Yu-kai, ZHAO Huan, NIU Jian-xin. Clonal Analysis of kfpSPL Gene and Its Promoter in Korla Fragrant Pear (Pyrus sinkiangensis) [J]. Xinjiang Agricultural Sciences, 2018, 55(6): 1017-1026. |
| [10] | HAO Xiao-yan, LI Jian-ping, Zumuremu Turxun, GAO Sheng-qi, HUANG Quan-sheng. Cloning and Bioinformatics Analysis of Salt-tolerance of PsaH Gene from Salicornia europaea [J]. Xinjiang Agricultural Sciences, 2017, 54(9): 1613-1620. |
| [11] | FENG Ya-ning;JIANG Zhi-qiang;MA Xiao-ling;LI Jiang-wei. Analysis of the Diversity of VHH Library from Bactrian Camel (Camelus bactrianus) with Bioinformatics [J]. , 2016, 53(8): 1533-1538. |
| [12] | GUO Chun-miao;YANG Bo;Tudi Maimaiti;LI Ning;TANG Ya-ping;GONG Peng;WANG Ji-xun. Studies on Polymorphism and Bioinformatics of 8 Xinjiang Almonds Self-incompatibility Gene (S-gene) [J]. , 2016, 53(6): 1067-1073. |
| [13] | . Bioinformatics Analysis of PSY in 21 Plant Species [J]. , 2015, 52(12): 2157-2165. |
| [14] | SU Li-bo;Ailizati Aili;LIU Liu;PENG Dan;YANG Yi;ZHANG Fu-chun. Cloning and Expression Analysis of Salt-stress Responsive Hc2a1 Gene from Halostachys caspica [J]. , 2014, 51(9): 1686-1691. |
| [15] | CHU Min;ZHANG Zhi-dong;WANG Wei;SONG Su-qing;Gu Mei-ying. Gene cloning and bioinformatics analysis of the cold-adapted amylase from Aeromonas sp. [J]. , 2014, 51(11): 2059-2065. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||