

新疆农业科学 ›› 2023, Vol. 60 ›› Issue (4): 857-864.DOI: 10.6048/j.issn.1001-4330.2023.04.009
陈果( ), 郝晓燕, 高升旗, 胡文冉, 赵准, 黄全生(
), 郝晓燕, 高升旗, 胡文冉, 赵准, 黄全生( )
)
收稿日期:2022-09-20
出版日期:2023-04-20
发布日期:2023-05-06
通信作者:
黄全生(1964-),男,乌鲁木齐人,研究员,博士,研究方向为作物抗逆分子生物学,(E-mail) 作者简介:陈果(1984-),男,四川仪陇人,副研究员,博士,研究方向为玉米分子生物学,(E-mail)252931257@qq.com
基金资助:
Chen Guo( ), Hao Xiaoyan, Gao Shengqi, Hu Wenran, Zhao Zhun, Huang Quansheng(
), Hao Xiaoyan, Gao Shengqi, Hu Wenran, Zhao Zhun, Huang Quansheng( )
)
Received:2022-09-20
Online:2023-04-20
Published:2023-05-06
Correspondence author:
HUANG Quansheng (1964-), Male, Urumqi, researcher, doctor, research direction: molecular biology of crop resistance, (E-mail) Supported by:摘要:
【目的】分析研究玉米钙依赖蛋白激酶全基因组鉴定及抗旱表达。【方法】使用NCBI Blast、DNAMAN和MotifScan等程序分析玉米钙依赖蛋白激酶CDPK的蛋白结构域和系统发育。采用Real-time RT-PCR方法分析CDPK基因在干旱胁迫处理玉米幼苗中的表达,评价CDPK基因对玉米干旱胁迫反应能力。【结果】鉴定出了在玉米中含有39个CDPK基因。将CDPK基因分为4个组。每个群体中的成员有共同的蛋白质基序和外显子及内含子结构,共同的进化起源。39个玉米CDPK基因根据它们的系统发育关系分组,并锚定在特定的玉米染色体上。多数CDPK基因在玉米干旱胁迫中的表达水平发生改变,CDPK基因参与对干旱胁迫的响应。【结论】从玉米基因组数据库中鉴定出了39个CDPK基因,大多数玉米CDPK基因在不同组织和不同发育阶段表现出不同的表达水平,CDPK基因在玉米发育中发挥不同的作用。多数CDPK基因在干旱胁迫下的表达量均发生变化。
中图分类号:
陈果, 郝晓燕, 高升旗, 胡文冉, 赵准, 黄全生. 玉米钙依赖蛋白激酶全基因组鉴定及抗旱表达分析[J]. 新疆农业科学, 2023, 60(4): 857-864.
Chen Guo, Hao Xiaoyan, Gao Shengqi, Hu Wenran, Zhao Zhun, Huang Quansheng. Genome-wide identification of the maize calcium-dependent protein kinase and drought expression analysis of the CDPK gene family in maize[J]. Xinjiang Agricultural Sciences, 2023, 60(4): 857-864.
| 名称 Name | 序列号 ID | CDs | 氨基酸 Amino acid | 分子量 MW (kDa) | EF手型数量 No. of EF Hands | N端序列 N-terminal | N端豆蔻酰化基元 N-Myristoylation |
|---|---|---|---|---|---|---|---|
| ZmCDPK1 | GRMZM2G028926_T01 | 1 827 | 608 | 66.4 | 4 | MGNTCVGP | No |
| ZmCDPK2 | GRMZM2G320506_T01 | 1 863 | 620 | 67.7 | 4 | MGNTCVGP | No |
| ZmCDPK3 | GRMZM2G025387_T01 | 1 593 | 530 | 59.9 | 4 | MGNRASRH | Yes |
| ZmCDPK4 | GRMZM2G035843_T01 | 1 527 | 508 | 56.5 | 4 | MQPDPSGN | No |
| ZmCDPK5 | GRMZM2G314396_T01 | 1 644 | 547 | 60.4 | 4 | MGNACSGA | Yes |
| ZmCDPK6 | GRMZM2G321239_T01 | 1 671 | 556 | 61.2 | 4 | MGNACGGA | Yes |
| ZmCDPK7 | GRMZM2G104125_T01 | 1 608 | 535 | 60.3 | 4 | MGNCCATP | Yes |
| ZmCDPK8 | AC210013.4_FGP014 | 1 617 | 538 | 60.4 | 4 | MGNCCAAP | Yes |
| ZmCDPK9 | GRMZM2G154489_T01 | 1 596 | 531 | 59.4 | 4 | MGQCCSRA | Yes |
| ZmCDPK10 | GRMZM2G030673_T01 | 1 626 | 541 | 60.5 | 4 | MGNCCRSP | No |
| ZmCDPK11 | GRMZM2G463464_T01 | 1 548 | 515 | 56.8 | 4 | MQPDPQGP | No |
| ZmCDPK12 | GRMZM2G047486_T01 | 1 533 | 510 | 60.2 | 4 | MQPDPSGN | No |
| ZmCDPK13 | GRMZM2G311220_T01 | 1 611 | 536 | 60.2 | 4 | MGNCCRSP | No |
| ZmCDPK14 | GRMZM2G028086_T01 | 1 620 | 539 | 60.8 | 4 | MGNCCVTP | Yes |
| ZmCDPK15 | GRMZM2G058305_T01 | 1 620 | 539 | 60.7 | 4 | MGGRASRH | Yes |
| ZmCDPK16 | GRMZM2G015703_T01 | 1 779 | 592 | 65.4 | 1 | MGLCHGKP | Yes |
| ZmCDPK17 | GRMZM2G167276_T01 | 1 533 | 510 | 56.2 | 4 | MGNCCPGS | Yes |
| ZmCDPK18 | GRMZM2G157068_T02 | 1 569 | 522 | 58.5 | 4 | MGACFSSA | Yes |
| ZmCDPK19 | GRMZM2G347047_T01 | 1 467 | 488 | 53.9 | 4 | MGGHQLHL | No |
| ZmCDPK20 | GRMZM2G121228_T01 | 1 743 | 580 | 63.7 | 4 | MGNTCVGP | No |
| ZmCDPK21 | GRMZM2G027351_T01 | 1 755 | 584 | 64.1 | 4 | MGNTCVGP | No |
| ZmCDPK22 | GRMZM2G347226_T01 | 1 548 | 515 | 56.8 | 4 | MQPDPQGS | No |
| ZmCDPK23 | GRMZM2G040743_T01 | 1 623 | 540 | 60.9 | 4 | MGNQCQNG | No |
| ZmCDPK24 | GRMZM2G080871_T02 | 1 536 | 511 | 57.0 | 3 | MGGCHSAI | No |
| ZmCDPK25 | GRMZM2G353957_T01 | 1 941 | 646 | 71.7 | 4 | MGNVCVGP | No |
| ZmCDPK26 | GRMZM2G081310_T01 | 1 689 | 562 | 61.9 | 4 | MGNTCGVT | No |
| ZmCDPK27 | GRMZM2G032852_T02 | 1 635 | 544 | 61.7 | 4 | MGNQCPNG | No |
| ZmCDPK28 | GRMZM2G365035_T01 | 1 539 | 512 | 57.5 | 4 | MGLCSSST | Yes |
| ZmCDPK29 | GRMZM2G112057_T01 | 1 620 | 539 | 61.0 | 4 | MGNCFTRK | Yes |
| ZmCDPK30 | GRMZM2G088361_T01 | 1 623 | 540 | 60.2 | 4 | MGNCCRSP | No |
| ZmCDPK31 | GRMZM5G856738_T03 | 1 575 | 524 | 59.5 | 4 | MGNRASRH | Yes |
| ZmCDPK32 | GRMZM2G099425_T01 | 1 620 | 539 | 60.8 | 4 | MGNCCVTP | Yes |
| ZmCDPK33 | GRMZM2G168706_T01 | 1 596 | 531 | 59.4 | 4 | MGQCCSRA | Yes |
| ZmCDPK34 | GRMZM2G340224_T01 | 1 842 | 613 | 67.5 | 4 | MRPSVSMI | Yes |
| ZmCDPK35 | GRMZM2G365815_T01 | 1 659 | 552 | 60.0 | 4 | MGQCCSKG | Yes |
| ZmCDPK36 | GRMZM2G472311_T01 | 1 746 | 581 | 63.4 | 4 | MGQCCSKG | Yes |
| ZmCDPK37 | AC203294.3_FGP001 | 1 395 | 464 | 50.9 | 4 | MGNCCPGS | No |
| ZmCDPK38 | GRMZM2G332660_T01 | 1 707 | 568 | 62.8 | 4 | MGGCYSAF | Yes |
| ZmCDPK39 | GRMZM2G097533_T01 | 1 317 | 438 | 48.5 | 1 | MGNCCVSR | Yes |
表1 玉米CDPKs的特征
Tab.1 Characteristics of CDPKs from maize
| 名称 Name | 序列号 ID | CDs | 氨基酸 Amino acid | 分子量 MW (kDa) | EF手型数量 No. of EF Hands | N端序列 N-terminal | N端豆蔻酰化基元 N-Myristoylation |
|---|---|---|---|---|---|---|---|
| ZmCDPK1 | GRMZM2G028926_T01 | 1 827 | 608 | 66.4 | 4 | MGNTCVGP | No |
| ZmCDPK2 | GRMZM2G320506_T01 | 1 863 | 620 | 67.7 | 4 | MGNTCVGP | No |
| ZmCDPK3 | GRMZM2G025387_T01 | 1 593 | 530 | 59.9 | 4 | MGNRASRH | Yes |
| ZmCDPK4 | GRMZM2G035843_T01 | 1 527 | 508 | 56.5 | 4 | MQPDPSGN | No |
| ZmCDPK5 | GRMZM2G314396_T01 | 1 644 | 547 | 60.4 | 4 | MGNACSGA | Yes |
| ZmCDPK6 | GRMZM2G321239_T01 | 1 671 | 556 | 61.2 | 4 | MGNACGGA | Yes |
| ZmCDPK7 | GRMZM2G104125_T01 | 1 608 | 535 | 60.3 | 4 | MGNCCATP | Yes |
| ZmCDPK8 | AC210013.4_FGP014 | 1 617 | 538 | 60.4 | 4 | MGNCCAAP | Yes |
| ZmCDPK9 | GRMZM2G154489_T01 | 1 596 | 531 | 59.4 | 4 | MGQCCSRA | Yes |
| ZmCDPK10 | GRMZM2G030673_T01 | 1 626 | 541 | 60.5 | 4 | MGNCCRSP | No |
| ZmCDPK11 | GRMZM2G463464_T01 | 1 548 | 515 | 56.8 | 4 | MQPDPQGP | No |
| ZmCDPK12 | GRMZM2G047486_T01 | 1 533 | 510 | 60.2 | 4 | MQPDPSGN | No |
| ZmCDPK13 | GRMZM2G311220_T01 | 1 611 | 536 | 60.2 | 4 | MGNCCRSP | No |
| ZmCDPK14 | GRMZM2G028086_T01 | 1 620 | 539 | 60.8 | 4 | MGNCCVTP | Yes |
| ZmCDPK15 | GRMZM2G058305_T01 | 1 620 | 539 | 60.7 | 4 | MGGRASRH | Yes |
| ZmCDPK16 | GRMZM2G015703_T01 | 1 779 | 592 | 65.4 | 1 | MGLCHGKP | Yes |
| ZmCDPK17 | GRMZM2G167276_T01 | 1 533 | 510 | 56.2 | 4 | MGNCCPGS | Yes |
| ZmCDPK18 | GRMZM2G157068_T02 | 1 569 | 522 | 58.5 | 4 | MGACFSSA | Yes |
| ZmCDPK19 | GRMZM2G347047_T01 | 1 467 | 488 | 53.9 | 4 | MGGHQLHL | No |
| ZmCDPK20 | GRMZM2G121228_T01 | 1 743 | 580 | 63.7 | 4 | MGNTCVGP | No |
| ZmCDPK21 | GRMZM2G027351_T01 | 1 755 | 584 | 64.1 | 4 | MGNTCVGP | No |
| ZmCDPK22 | GRMZM2G347226_T01 | 1 548 | 515 | 56.8 | 4 | MQPDPQGS | No |
| ZmCDPK23 | GRMZM2G040743_T01 | 1 623 | 540 | 60.9 | 4 | MGNQCQNG | No |
| ZmCDPK24 | GRMZM2G080871_T02 | 1 536 | 511 | 57.0 | 3 | MGGCHSAI | No |
| ZmCDPK25 | GRMZM2G353957_T01 | 1 941 | 646 | 71.7 | 4 | MGNVCVGP | No |
| ZmCDPK26 | GRMZM2G081310_T01 | 1 689 | 562 | 61.9 | 4 | MGNTCGVT | No |
| ZmCDPK27 | GRMZM2G032852_T02 | 1 635 | 544 | 61.7 | 4 | MGNQCPNG | No |
| ZmCDPK28 | GRMZM2G365035_T01 | 1 539 | 512 | 57.5 | 4 | MGLCSSST | Yes |
| ZmCDPK29 | GRMZM2G112057_T01 | 1 620 | 539 | 61.0 | 4 | MGNCFTRK | Yes |
| ZmCDPK30 | GRMZM2G088361_T01 | 1 623 | 540 | 60.2 | 4 | MGNCCRSP | No |
| ZmCDPK31 | GRMZM5G856738_T03 | 1 575 | 524 | 59.5 | 4 | MGNRASRH | Yes |
| ZmCDPK32 | GRMZM2G099425_T01 | 1 620 | 539 | 60.8 | 4 | MGNCCVTP | Yes |
| ZmCDPK33 | GRMZM2G168706_T01 | 1 596 | 531 | 59.4 | 4 | MGQCCSRA | Yes |
| ZmCDPK34 | GRMZM2G340224_T01 | 1 842 | 613 | 67.5 | 4 | MRPSVSMI | Yes |
| ZmCDPK35 | GRMZM2G365815_T01 | 1 659 | 552 | 60.0 | 4 | MGQCCSKG | Yes |
| ZmCDPK36 | GRMZM2G472311_T01 | 1 746 | 581 | 63.4 | 4 | MGQCCSKG | Yes |
| ZmCDPK37 | AC203294.3_FGP001 | 1 395 | 464 | 50.9 | 4 | MGNCCPGS | No |
| ZmCDPK38 | GRMZM2G332660_T01 | 1 707 | 568 | 62.8 | 4 | MGGCYSAF | Yes |
| ZmCDPK39 | GRMZM2G097533_T01 | 1 317 | 438 | 48.5 | 1 | MGNCCVSR | Yes |
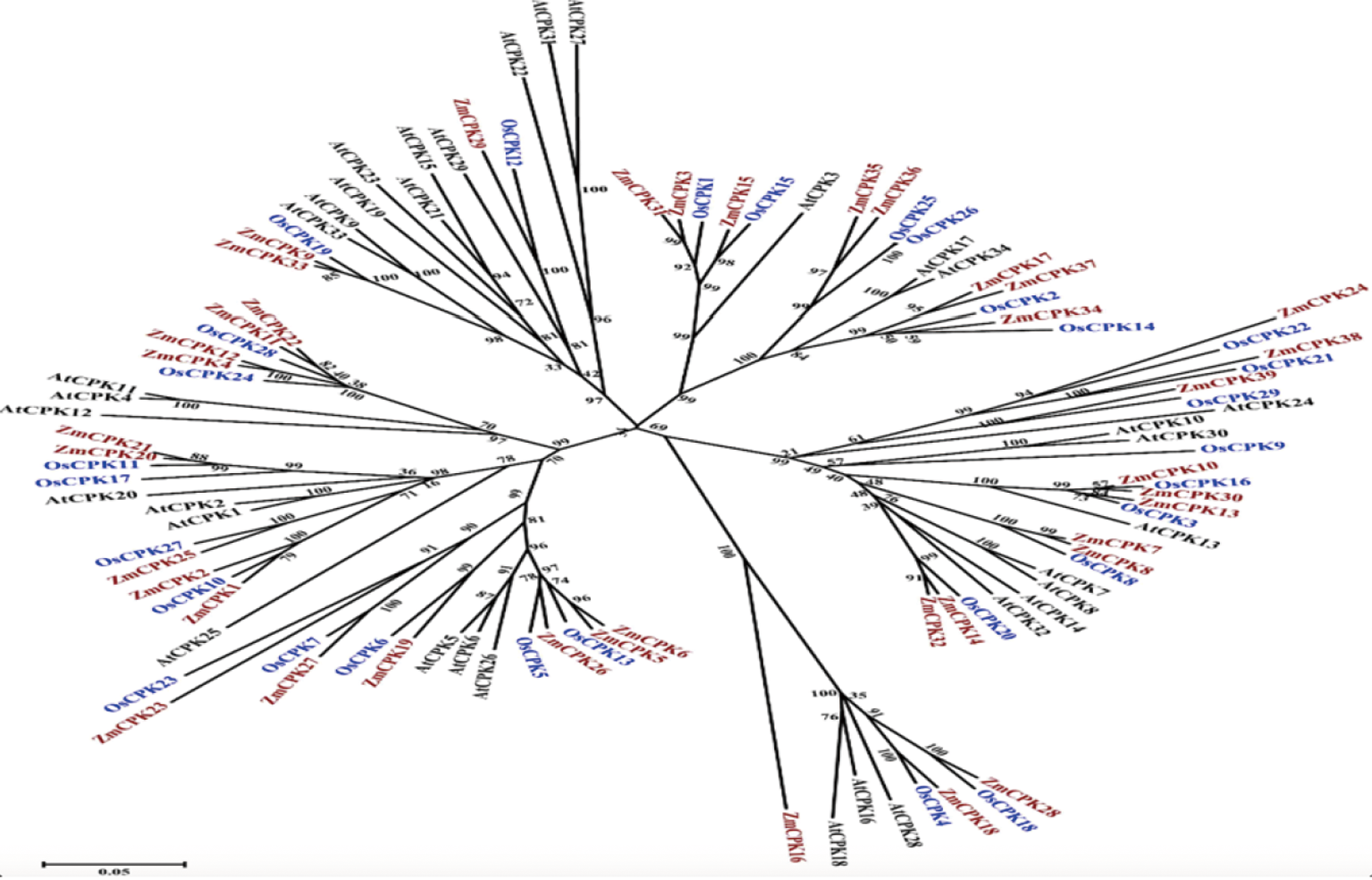

图1 玉米、水稻和拟南芥CDPKs的系统发育树 注:利用39个玉米、29个水稻和34个拟南芥的CDPK蛋白全长序列,使用MEGA5.0程序建立玉米CDPK基因的系统发育树。棕色代表玉米CDPK基因;蓝色代表水稻CDPK基因,黑色代表拟南芥CDPK基因
Fig.1 Phylogenetic tree of CDPKs from maize, rice and Arabidopsis Note:Neighbor-joining tree was created using MEGA5.0 program with 1,000 bootstrap using full length sequences of 39 maize, 29 rice, and 34 Arabidopsis CDPK proteins. Brown, maize orthologs; blue, rice orthologs, black, Arabidopsis orthologs
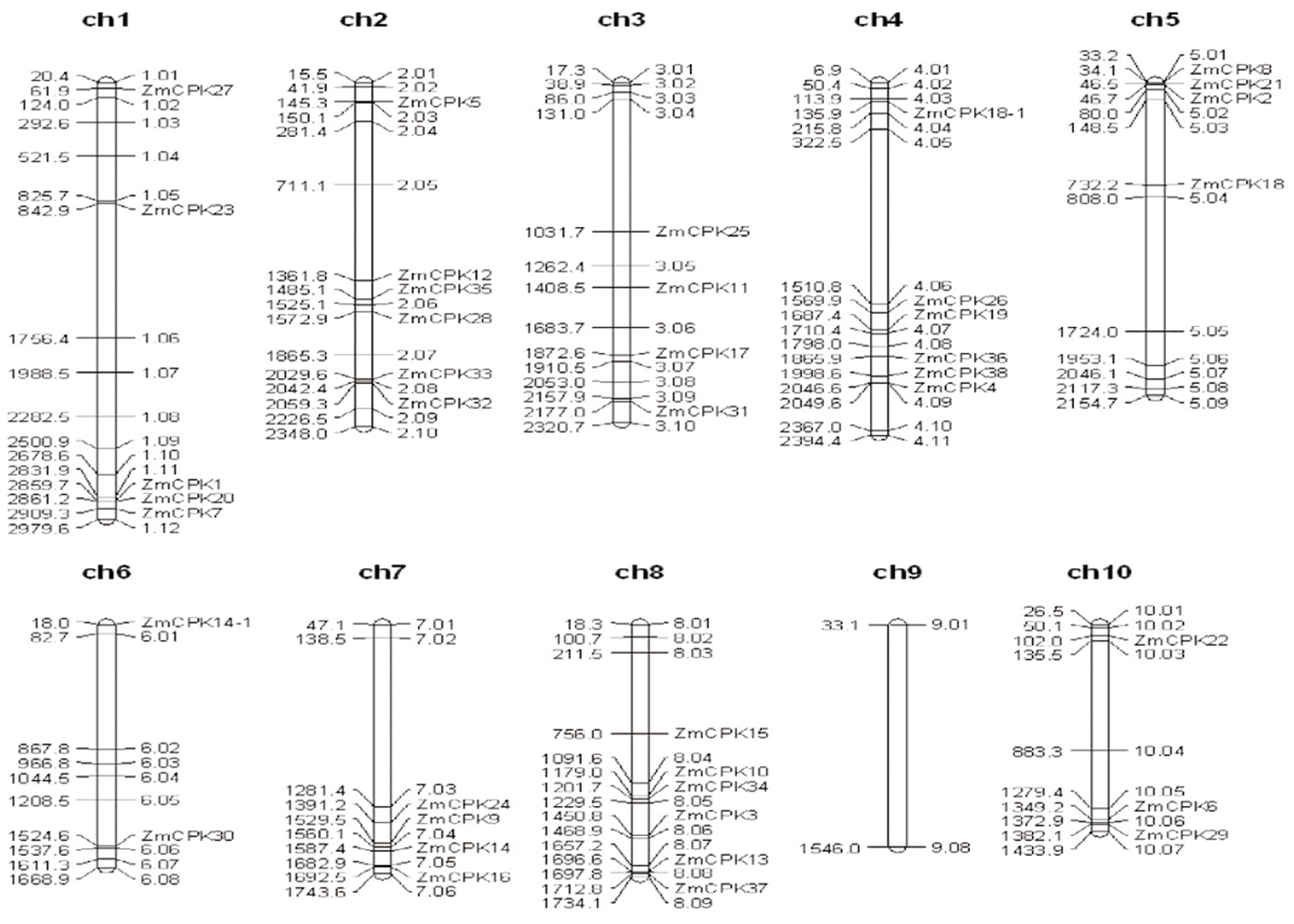
图2 玉米基因组CDPK基因的染色体分布 注:染色体编号在每个染色体表示法的顶部标明
Fig.2 Chromosomal distributions of CDPK genes in the maize genome Note:The chromosome number is indicated at the top of each chromosome representation
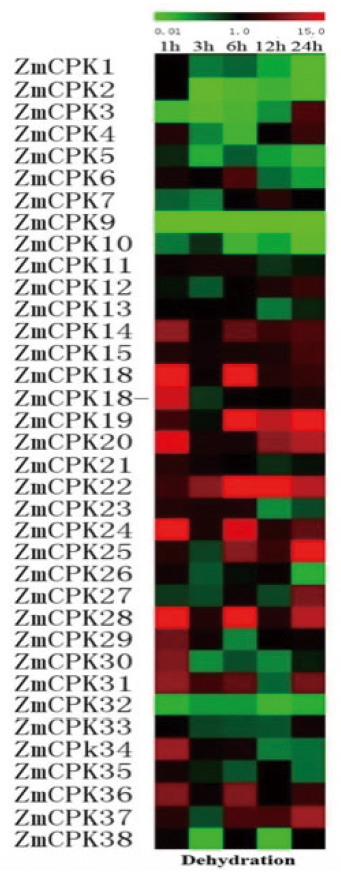
图3 实时定量RT-PCR分析39个CDPK基因在不同时间干旱胁迫下的表达 注:图中所示数字表示相对信号强度值,采用了分层聚类进行数据分析
Fig.3 Expression analysis of 39 CDPK genes under drought stress for various times as indicated by quantitative real-time RT-PCR analysis Note:The scale representing the relative signal intensity values is shown above. Hierarchical clustering was played in data analysis
| [1] | 梁晓玲, 刘文欣, 阿布来提·阿布拉, 等. 干旱胁迫对玉米杂交种产量及穗部性状的影响[J]. 玉米科学, 2021, 29(2):75-80. |
| Liang Xiaoling, Liu Wenxin, Abulaiti Abra, et al. Influence of drought stress on yield and ear traits of maize hybrids[J]. Journal of Maize Sciences, 2021, 29(2):75-80. | |
| [2] | Hu H, Xiong L. Genetic engineering and breeding of drought-resistant crops[J]. Annu Rev Plant Biol, 2014,(65):715-741. |
| [3] |
Tuteja N, Mahajan S. Calcium signaling network in plants: an overview[J]. Plant Signal Behav, 2007, (2):79-85.
DOI PMID |
| [4] | Harper JF, Harmon A. Plants symbiosis and parasites: a calcium signaling connection[J]. Nature Rev, 2005, (6):555-566. |
| [5] | Lourido S, Shuman J, Zhang C, et al. Calcium dependent protein kinase 1 is an essential regulator of exocytosis in Toxoplasma[J]. Nature, 2010, (465):359-362. |
| [6] |
Klimecka M, Muszynska G. Structure and functions of plant calcium dependent protein kinases[J]. Acta Biochim Pol, 2007, 54:219-233.
PMID |
| [7] |
Romeis T, Ludwig AA, Martin R, et al. Calcium-dependent protein kinases play an essential role in a plant defence response[J]. EMBO J, 2001, (20):5556-5567.
DOI PMID |
| [8] | Asano T, Hayashi N, Kikuchi S, et al. CDPK-mediated abiotic stress signaling[J]. Plant Signal Behav, 2012, (7):1-5. |
| [9] | Lee J, Rudd JJ. Calcium-dependent protein kinases: versatile plant signaling components necessary for pathogen defence[J]. Trends Plant Sci, 2002, (7):97-98. |
| [10] | Ludwig AA, Romeis T, Jones JD. CDPK-mediated signalling pathways: specificity and cross-talk[J]. J Exp Bot, 2004, (55):181-188. |
| [11] | Zhu SY, Yu XC, Wang XJ, et al. Two calcium-dependent protein kinases CPK4 and CPK11 regulate abscisic acid signal transduction in Arabidopsis[J]. Plant Cell, 2007, (19):3019-3036. |
| [12] | Mori IC, Murata Y, Yang Y, et al. CDPKs CPK6 and CPK3 function in ABA regulation of guard cell S-type anion- and Ca2+-permeable channels and stomatal closure[J]. PLoS Biol, 2006, (4): e327. |
| [13] | Munemasa S, Hossain MA, Nakamura Y, et al. The Arabidopsis calcium-dependent protein kinase CPK6 functions as a positive regulator of methyl jasmonate signaling in guard cells[J]. Plant Physiol, 2011, (155):553-561. |
| [14] | Ma SY, Wu WH. AtCPK23 functions in Arabidopsis responses to drought and salt stresses[J]. Plant Mol Biol, 2007, (65):511-518. |
| [15] | Franz S, Ehlert B, Liese A, et al. Calcium-dependent protein kinase CPK21 functions in abiotic stress response in Arabidopsis thaliana[J]. Mol Plant, 2011, (4):83-96. |
| [16] |
Choi HI, Park HJ, Park JH, et al. Arabidopsis calcium-dependent protein kinase AtCPK32 interacts with ABF4 a transcriptional regulator of abscisic acid-responsive gene expression and modulates its activity[J]. Plant Physiol, 2005, 139:1750-1761.
DOI URL |
| [17] | Boudsocq M, Willmann MR, McCormack M, et al. Differential innate immune signaling via Ca2+sensor protein kinases[J]. Nature, 2010, (464):418-422. |
| [18] | Saijo Y, Hata S, Kyozuka J, et al. Over-expression of a single Ca2+dependent protein kinase confers both cold and salt/drought tolerance on rice plants[J]. Plant J, 2000, (23):319-327. |
| [19] | Asano T, Hakata M, Nakamura H, et al. Functional characterisation of OsCPK21 a calcium-dependent protein kinase that confers salt tolerance in rice[J]. Plant Mol Biol, 2011, (75):179-191. |
| [20] | Asano T, Hayashi N, Kobayashi M, et al. A rice calcium-dependent protein kinase OsCPK12 oppositely modulates salt-stress tolerance and blast disease resistance[J]. Plant J, 2012, (69):26-36. |
| [21] | Cheng SH, Willmann MR, Chen HC, et al. Calcium signaling through protein kinases: The Arabidopsis calcium-dependent protein kinase gene family[J]. Plant Physiol, 2002, (129):469-485. |
| [22] | Asano T, Tanaka N, Yang G, et al. Genome-wide identification of the rice calcium-dependent protein kinase and its closely related kinase gene families: comprehensive analysis of the CDPKs gene family in rice[J]. Plant Cell Physiol, 2005, (46):356-366 |
| [23] | Li AL, Zhu YF, Tan XM, et al. Evolutionary and functional study of the CDPK gene family in wheat (Triticum aestivum L)[J]. Plant Mol Biol, 2008, (66):429-443. |
| [24] | Yang DH, Hettenhausen C, Baldwin IT, et al. Nicotiana attenuata calcium-dependent protein kinases CDPK4 and CDPK5 strongly downregulate wound- and herbivory-induced jasmonic acid accumulations[J]. Plant Physiol, 2012, (159):1561-1607. |
| [25] | Giammaria V, Grandellis C, Bachmann S, et al. StCDPK2 expression and activity reveal a highly responsive potato calcium-dependent protein kinase involved in light signaling[J]. Planta, 2011, (233):593-609. |
| [26] | Chang WJ, Su HS, Li WJ, et al. Expression profiling of a novel calcium-dependent protein kinase gene LeCPK2 from tomato (Solanum lycopersicum) under heat and pathogen-related hormones[J]. Biosci Biotech Bioch, 2009, (73):2427-2431. |
| [27] | Berberich T, Kusano T. Cycloheximide induces a subset of low temperature-inducible genes in maize[J]. Mol Gen Genet, 1997, (254):275-283. |
| [28] | Saijo Y, Hata S, Sheen J, et al. cDNA cloning and prokaryotic expression of maize calcium-dependent protein kinases[J]. Biochim Biophys Acta, 1997, (1350):109-114. |
| [29] | Murillo I, Jaeck E, Cordero MJ, et al. Transcriptional activation of a maize calcium-dependent protein kinase gene in response to fungal elicitors and infection[J]. Plant Mol Biol, 2001, (45):145-158. |
| [30] | Klimecka M, Szczegielniak J, Godecka L, et al. Regulation of wound-responsive calcium dependent protein kinase from maize (ZmCPK11) by phosphatidic acid[J]. Acta Biochim Pol, 2011, (58):589-595. |
| [31] | Schnable PS, Ware D, Fulton RS, et al. The B73 maize genome: complexity diversity and dynamics[J]. Science, 2009, (326):1112-1115. |
| [32] | Matschi S, Werner S, Schulze WX, et al. Function of calcium-dependent protein kinase CPK28 of Arabidopsis thaliana in plant stem elongation and vascular development[J]. Plant J, 2013, (73):883-896. |
| [1] | 王晓雨, 王小平, 史文宇, 刘美艳, 马健, 郭云鹏, 宋瑞欣, 王清涛. 拔节期冬小麦光合特性、干物质积累和产量对干旱胁迫的响应[J]. 新疆农业科学, 2023, 60(9): 2163-2172. |
| [2] | 向莉, 王仙, 董裕生, 郭小玲, 方伏荣, 陈智军, 马艳明, 苗雨. 外源丁酸对干旱胁迫下大麦产量及品质的影响[J]. 新疆农业科学, 2023, 60(9): 2173-2181. |
| [3] | 米尔扎提·木塔力甫, 石秀楠, 柏军兵, 祖拜代·阿布都克日木, 吾勒加勒哈斯·阿扎提, 石书兵. 不同脱绒方式及PEG胁迫下对棉花种子活力及幼苗性状的影响[J]. 新疆农业科学, 2023, 60(7): 1561-1568. |
| [4] | 王挺, 张力, 张凡凡, 黄嵘峥, 李肖, 张玉琳, 陈永成, 赵建涛, 马春晖. 适合青贮的玉米品种生产性能筛选及营养价值评价[J]. 新疆农业科学, 2023, 60(7): 1596-1605. |
| [5] | 陈占辉, 孙强, 任姣姣, 黄博文, 许加波, 杨杰, 吴鹏昊. 玉米叶宽的QTL定位及全基因组选择分析[J]. 新疆农业科学, 2023, 60(7): 1606-1613. |
| [6] | 蓝陈仪航, 姚禹博, 周俊祥, 付开赟, 丁新华, 尹晓辉, 刘文, 王娜, 郭文超, 邓建宇. 亚洲玉米螟性信息素鉴定及其应用研究进展[J]. 新疆农业科学, 2023, 60(7): 1614-1622. |
| [7] | 曲可佳, 时晓磊, 张恒, 王兴州, 耿洪伟, 丁孙磊, 张金波, 严勇亮. PEG处理下引进春小麦品种苗期抗旱性评价[J]. 新疆农业科学, 2023, 60(6): 1363-1371. |
| [8] | 邵盘霞, 赵准, 邵武奎, 郝晓燕, 高升旗, 李建平, 胡文冉, 黄全生. 玉米ZmCDPK22基因在干旱胁迫下的表达分析[J]. 新疆农业科学, 2023, 60(6): 1372-1378. |
| [9] | 陆晏天, 桑志勤, 徐灿, 张力, 夏春兰, 王友德, 李伟, 陈树宾. 历年新疆和宁夏审定玉米品种主要性状的演化及品种审定现状分析[J]. 新疆农业科学, 2023, 60(6): 1379-1388. |
| [10] | 耿翡翡, 孟超敏, 卿桂霞, 周佳敏, 张富厚, 刘逢举. 陆地棉磷高效基因GhMYB4的克隆与表达分析[J]. 新疆农业科学, 2023, 60(6): 1406-1412. |
| [11] | 汤东, 安玉光, 程平, 李宏, 杨建军, 王凯. 天山北坡前山带典型灌木光合特性对干旱胁迫的响应[J]. 新疆农业科学, 2023, 60(6): 1531-1539. |
| [12] | 杨明花, 刘强, 廖必勇, 彭云承, 布阿依夏木·那曼提, 达吾来·杰克山. 不完全双列杂交玉米组合抗倒伏综合评价[J]. 新疆农业科学, 2023, 60(4): 832-840. |
| [13] | 周广威, 韩登旭, 朱琦, 张少民. 耐低磷新疆春玉米基因型筛选及其磷效率[J]. 新疆农业科学, 2023, 60(4): 847-856. |
| [14] | 董秀丽, 韩登旭, 杨杰, 阿布来提·阿布拉, 戴爱梅, 李俊杰, 王业建, 刘俊, 郗浩江, 梁晓玲, 李铭东. 密植条件下玉米主要农艺性状的综合性分析[J]. 新疆农业科学, 2023, 60(4): 865-871. |
| [15] | 侯良忠, 郭同军, 张俊瑜, 苏玲玲, 古再丽努尔·艾麦提, 温小燕, 朱晓芳, 杨建中, 王文奇. 不同比例籽用西葫芦皮瓤与玉米秸秆混贮效果评价[J]. 新疆农业科学, 2023, 60(3): 567-573. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||