

Xinjiang Agricultural Sciences ›› 2023, Vol. 60 ›› Issue (2): 389-398.DOI: 10.6048/j.issn.1001-4330.2023.02.016
• Horticultural Special Local Products·Plant Protection·Microbes·Soil Fertilizer· Water Saving Irrigation • Previous Articles Next Articles
SONG Jindi( ), LIU Jun(
), LIU Jun( ), SUN Yufang, Youlituzi Naibi, CHEN Baoqiang, XIE Bingbing
), SUN Yufang, Youlituzi Naibi, CHEN Baoqiang, XIE Bingbing
Received:2022-06-22
Online:2023-02-20
Published:2023-03-31
Correspondence author:
LIU Jun (1970-), female, born in Urumqi, Xinjiang, Associate professor, postgraduate supervisor, research interest: plant pathogenic bacteriology, (E-mail)liujem@126.com
Supported by:
宋金迪( ), 刘君(
), 刘君( ), 孙玉芳, 优丽图孜·乃比, 陈宝强, 颉兵兵
), 孙玉芳, 优丽图孜·乃比, 陈宝强, 颉兵兵
通讯作者:
刘君(1970-),女,新疆乌鲁木齐人,副教授,硕士研究生导师,研究方向为植物病原细菌学,(E-mail)liujem@126.com
作者简介:宋金迪(1994-),男,河南洛阳人,硕士研究生,研究方向为植物病原细菌学,(E-mail)sjddog@126.com
基金资助:CLC Number:
SONG Jindi, LIU Jun, SUN Yufang, Youlituzi Naibi, CHEN Baoqiang, XIE Bingbing. Transcriptome Analysis of Acidovorax citrulli against Copper Stress[J]. Xinjiang Agricultural Sciences, 2023, 60(2): 389-398.
宋金迪, 刘君, 孙玉芳, 优丽图孜·乃比, 陈宝强, 颉兵兵. 铜胁迫下的西瓜食酸菌转录组分析[J]. 新疆农业科学, 2023, 60(2): 389-398.
Add to citation manager EndNote|Ris|BibTeX
URL: http://www.xjnykx.com/EN/10.6048/j.issn.1001-4330.2023.02.016
| 基因ID Gene ID | 本地数据库基因ID Local database gene ID | 正向引物(5'~3') Forward primer | 反向引物(5'~3') Reverse primer |
|---|---|---|---|
| NMDCN0000J7Q | GE00642 | GCGTGACCGCCATCCAG | CCTTGCGGGCGATGATG |
| NMDCN0000J7N | GE00282 | AACAACTACCAGGTGGGACT | GGTGAGCTGCACGGATT |
| NMDCN0000J7P | GE01538 | AGCCTCTTGCCATTGCC | CACCACCTGGGTATCGC |
| NMDCN0000J7O | GE02901 | CCCTCCGCCAGAACAAT | TCCACCGACACGATGATG |
| NMDCN0000J7M | GE00281 | GCGCTACAACGGCTACC | GCCAGGCAGAGGAACAC |
| NMDCN0000J7L | GE01342 | TGCCGTGACGACTGCTT | GCGTATTGTTGGCCTTGT |
| NMDCN0000J7K | GE04032 | AACAACTACCAGGTGGGACT | GGTGAGCTGCACGGATT |
| NMDCN0000J7J | GE01848 | CGCCTATCCGTCCAAGC | AGCACCGTGTAGCCGTC |
| NMDCN0000J7I | GE02981 | CGCCGTCCCTCCCGTTT | GCCAGGATCGTCCCCAG |
| NMDCN0000J7H | GE01751 | GTCAACCTCTGGGGCTACA | AACTCGTACACGAAGGTCTTG |
| NMDCN0000J7G | GE01955 | GAACCTGGAAGTCATCATCG | CCTTGGTCTTGTTGTCGG |
| NMDCN0000Q79 | GE04262 | GGCTGGCTGGGTCATAC | ATTGTAGATTTCGCCGTTG |
| NMDCN0000Q7A | GE02797 | CACAAGACTGTTTCTGTTCCAAG | GCACGTATGTCCGTCACCT |
| NMDCN0000Q7B | GE00664 | CGTCCCGTATCGCCGTCA | GCGTGATGCCCGTGAGC |
| NMDCN0000Q7C | GE01086 | GCTGCTGTCGGAAATGG | TTGTTGGTGCGGAAGTTAC |
| rpoB | - | GCGACAGCGTGCTCAAAGTG | GGCCTTCGTTGGTGCGTTTCT |
Table 1 Primers used for validation qPCR of transcriptome data
| 基因ID Gene ID | 本地数据库基因ID Local database gene ID | 正向引物(5'~3') Forward primer | 反向引物(5'~3') Reverse primer |
|---|---|---|---|
| NMDCN0000J7Q | GE00642 | GCGTGACCGCCATCCAG | CCTTGCGGGCGATGATG |
| NMDCN0000J7N | GE00282 | AACAACTACCAGGTGGGACT | GGTGAGCTGCACGGATT |
| NMDCN0000J7P | GE01538 | AGCCTCTTGCCATTGCC | CACCACCTGGGTATCGC |
| NMDCN0000J7O | GE02901 | CCCTCCGCCAGAACAAT | TCCACCGACACGATGATG |
| NMDCN0000J7M | GE00281 | GCGCTACAACGGCTACC | GCCAGGCAGAGGAACAC |
| NMDCN0000J7L | GE01342 | TGCCGTGACGACTGCTT | GCGTATTGTTGGCCTTGT |
| NMDCN0000J7K | GE04032 | AACAACTACCAGGTGGGACT | GGTGAGCTGCACGGATT |
| NMDCN0000J7J | GE01848 | CGCCTATCCGTCCAAGC | AGCACCGTGTAGCCGTC |
| NMDCN0000J7I | GE02981 | CGCCGTCCCTCCCGTTT | GCCAGGATCGTCCCCAG |
| NMDCN0000J7H | GE01751 | GTCAACCTCTGGGGCTACA | AACTCGTACACGAAGGTCTTG |
| NMDCN0000J7G | GE01955 | GAACCTGGAAGTCATCATCG | CCTTGGTCTTGTTGTCGG |
| NMDCN0000Q79 | GE04262 | GGCTGGCTGGGTCATAC | ATTGTAGATTTCGCCGTTG |
| NMDCN0000Q7A | GE02797 | CACAAGACTGTTTCTGTTCCAAG | GCACGTATGTCCGTCACCT |
| NMDCN0000Q7B | GE00664 | CGTCCCGTATCGCCGTCA | GCGTGATGCCCGTGAGC |
| NMDCN0000Q7C | GE01086 | GCTGCTGTCGGAAATGG | TTGTTGGTGCGGAAGTTAC |
| rpoB | - | GCGACAGCGTGCTCAAAGTG | GGCCTTCGTTGGTGCGTTTCT |
| 样品名 Sample | Reads总数 Total reads | 碱基总数 Bases(bp) | N (%) | Q20 (%) | Q30 (%) |
|---|---|---|---|---|---|
| CK1 | 29 540 900 | 4 460 675 900 | 0.04 | 96.90 | 92.01 |
| CK2 | 31 451 022 | 4 749 104 322 | 0.04 | 96.80 | 91.82 |
| CK3 | 27 676 892 | 4 179 210 692 | 0.02 | 97.09 | 92.40 |
| T1 | 29 357 120 | 4 432 925 120 | 0.02 | 96.93 | 92.02 |
| T2 | 27 027 648 | 4 081 174 848 | 0.01 | 96.99 | 92.14 |
| T3 | 27 104 738 | 4 092 815 438 | 0.02 | 97.04 | 92.24 |
Table 2 Base mass statistics of the transcript of Acidovorax citrulli FC440 under copper stress
| 样品名 Sample | Reads总数 Total reads | 碱基总数 Bases(bp) | N (%) | Q20 (%) | Q30 (%) |
|---|---|---|---|---|---|
| CK1 | 29 540 900 | 4 460 675 900 | 0.04 | 96.90 | 92.01 |
| CK2 | 31 451 022 | 4 749 104 322 | 0.04 | 96.80 | 91.82 |
| CK3 | 27 676 892 | 4 179 210 692 | 0.02 | 97.09 | 92.40 |
| T1 | 29 357 120 | 4 432 925 120 | 0.02 | 96.93 | 92.02 |
| T2 | 27 027 648 | 4 081 174 848 | 0.01 | 96.99 | 92.14 |
| T3 | 27 104 738 | 4 092 815 438 | 0.02 | 97.04 | 92.24 |
| 样品名 Sample | 可用读取 Useful Reads | 总映射读取 Total Mapped Reads | 占比 Proportion (%) | 单映射读取 Uniquely Mapped Reads | 占比 Proportion (%) | 多映射读取 Multiple Mapped Reads | 占比 Proportion (%) |
|---|---|---|---|---|---|---|---|
| CK1 | 29 395 754 | 29 311 584 | 99.71 | 28 350 407 | 96.72 | 961 177 | 3.28 |
| CK2 | 31 279 164 | 31 182 372 | 99.69 | 30 036 408 | 96.32 | 1 145 964 | 3.68 |
| CK3 | 27 553 118 | 27 468 703 | 99.69 | 26 767 147 | 97.45 | 701 556 | 2.55 |
| T1 | 29 214 302 | 29 096 642 | 99.60 | 28 357 073 | 97.46 | 739 569 | 2.54 |
| T2 | 26 898 484 | 26 777 059 | 99.55 | 26 086 501 | 97.42 | 690 558 | 2.58 |
| T3 | 26 987 212 | 26 892 876 | 99.65 | 26 195 932 | 97.41 | 696 944 | 2.59 |
Table 3 Reference genome comparison of Acidovorax citrulli FC440
| 样品名 Sample | 可用读取 Useful Reads | 总映射读取 Total Mapped Reads | 占比 Proportion (%) | 单映射读取 Uniquely Mapped Reads | 占比 Proportion (%) | 多映射读取 Multiple Mapped Reads | 占比 Proportion (%) |
|---|---|---|---|---|---|---|---|
| CK1 | 29 395 754 | 29 311 584 | 99.71 | 28 350 407 | 96.72 | 961 177 | 3.28 |
| CK2 | 31 279 164 | 31 182 372 | 99.69 | 30 036 408 | 96.32 | 1 145 964 | 3.68 |
| CK3 | 27 553 118 | 27 468 703 | 99.69 | 26 767 147 | 97.45 | 701 556 | 2.55 |
| T1 | 29 214 302 | 29 096 642 | 99.60 | 28 357 073 | 97.46 | 739 569 | 2.54 |
| T2 | 26 898 484 | 26 777 059 | 99.55 | 26 086 501 | 97.42 | 690 558 | 2.58 |
| T3 | 26 987 212 | 26 892 876 | 99.65 | 26 195 932 | 97.41 | 696 944 | 2.59 |
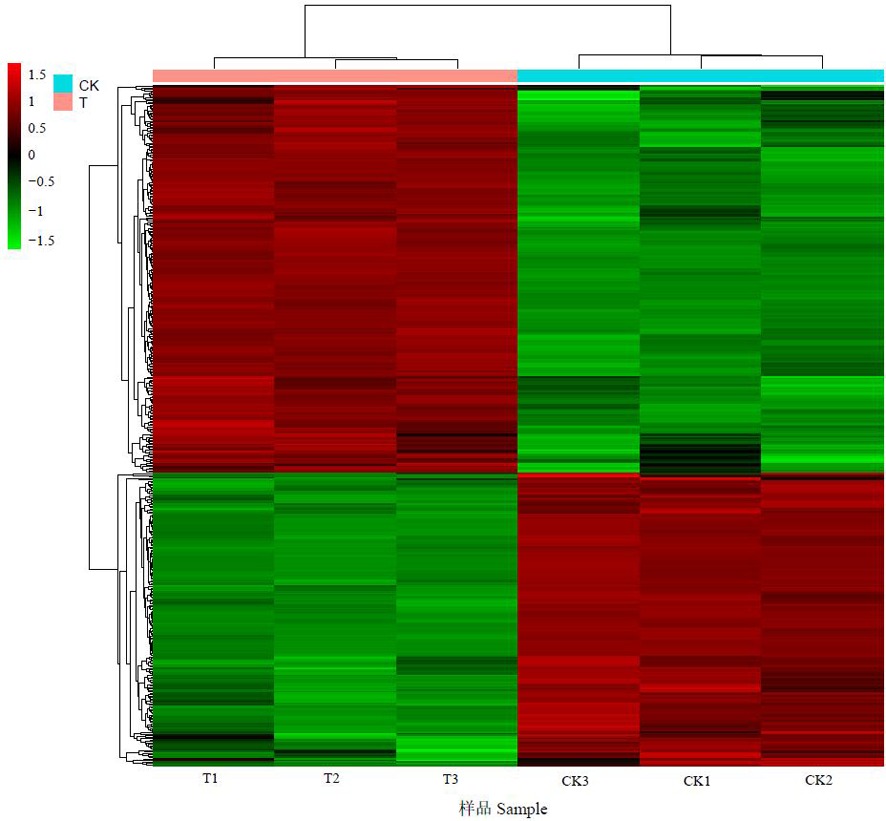
Fig.3 Clustering heat map of differentially expressed genes between samples of Acidovorax citrulli FC440 under copper stress Note: Horizontal represents the gene, each column is a sample, red represents the high expressed gene, green represents the low expressed gene

Fig.7 The KEGG pathway significantly enriched bubble diagram Note: The ordinate is KEGG Pathway entry;The horizontal axis is Richfactor.The size of the dots in the figure indicates the number of differential genes annotated to this pathway, and the color indicates the significant FDR value of this pathway
| [1] |
Schaad N W, Postnikova E, Sechler A, et al. Reclassification of subspecies of Acidovorax avenae as A.Avenae (Manns 1905) emend., A.cattleyae (Pavarino, 1911) comb.nov., A.citrulli Schaad et al., 1978) comb.nov., and proposal of A.oryzae sp.nov[J]. Systematic and Applied Microbiology, 2008, 31(6-8): 434-446.
DOI URL |
| [2] |
Bahar O, Burdman S. Bacterial fruit blotch: A threat to the cucurbit industry[J]. Israel Journal of Plant Sciences, 2010, 58(1): 19-31.
DOI URL |
| [3] | Latin R X. Bacterial Fruit Blotch of Watermelon in Indiana[J]. Plant Disease, 1990, 74(4): 331-342. |
| [4] | 赵文龙. 瓜类细菌性果斑病菌抗铜性研究[D]. 北京: 中国农业科学院, 2013. |
| ZHAO Wenlong. Analysis of copper resistance in Acidovorax citrulli[D]. Beijing: Chinese Academy of Agricultural Sciences, 2013. | |
| [5] | 李强. 瓜类果斑病菌多铜氧化酶基因cueO的功能研究[D]. 呼和浩特: 内蒙古农业大学, 2014. |
| LI Qiang. Functional Analysis of cueO gene in the bacterial fruit blotch of watermelon[D]. Hohhot: Inner Mongolia Agricultural University, 2014. | |
| [6] | 刘星, 王希东, 刘君. 西瓜食酸菌RND蛋白家族外排转运体cusB基因抗铜功能研究[J]. 微生物学通报, 2016, 43(1): 97-106. |
| LIU Xing, WANG Xidong, LIU Jun. Functional analysis of a RND family effiux transporter component-cusB gene associated with copper resistance in Acidovorax citrulli[J]. Microbiology China, 2016, 43(1): 97-106. | |
| [7] | 颉兵兵, 刘君, 优丽图孜·乃比, 等. 西瓜食酸菌抗铜基因cueR的生物信息学分析及功能验证[J]. 微生物学通报, 2020, 47(5): 1534-1543. |
| XIE Bingbing, LIU Jun, Youlituzi Naibi et al. Informatics analysis and functional verification of copper resistance gene cueR in Acidovorax citrulli[J]. Microbiology China, 2020, 47(5): 1534-1543. | |
| [8] | Martin M. Cut adapt removes adapter sequences from high-throughput sequencing reads[J]. Embnet Journal, 2011, 17(1): 10-12. |
| [9] | Eckshtain-Levi N, Shkedy D, Gershovits M, et al. Insights from the genome sequence of Acidovorax citrulli M6, a group I strain of the causal agent of bacterial fruit blotch of cucurbits[J]. Frontiers in microbiology, 2016, 7(1): 430-441. |
| [10] |
Mortazavi A, Williams B A, McCue K, et al. Mapping and quantifying mammalian transcriptomes by RNA-Seq[J]. Nature Methods, 2008, 5(7): 621-628.
DOI PMID |
| [11] | 王立坤. RNA-seq数据的处理与应用[D]. 长春: 吉林大学, 2012. |
| WANG Likun. Processing and application of RNA-seq data[D]. Changchun: Jilin University. | |
| [12] | Wang T, Sun B, Yang Y, et al. Genome sequence of Acidovorax citrulli group 1 Strain pslb65 causing bacterial fruit blotch of melons[J]. Genome Announcements, 2015, 3(2): 315-327. |
| [13] | Anders S, Huber W. Differential expression analysis for sequence count data[J]. Nature Precedings, 2010, 11: R106. |
| [14] |
Nanda M, Kumar V, Sharma D K. Multimetal tolerance mechanisms in bacteria: The resistance strategies acquired by bacteria that can be exploited to ‘clean-up’ heavy metal contaminants from water[J]. Aquatic Toxicology, 2019, 212: 1-10.
DOI URL |
| [15] | Nies D H. Efflux-mediated heavy metal resistance in prokaryotes[J]. FEMS Microbiology Reviews, 2003, 781(2-3): 313-339. |
| [16] | 黄成文. 西瓜食酸菌抗铜相关基因tolC和gntR的鉴定与功能分析[D]. 乌鲁木齐: 新疆农业大学, 2018. |
| HUANG Chengwen. Identification and functional analysis of copper resistance genes tolC and gntR in Acidovorax citrulli[D]. Urumqi: Xinjiang Agricultural University, 2018. | |
| [17] |
Macomber L, Hausinger R P. Mechanisms of nickel toxicity in microorganisms[J]. Metallomics, 2011, 3(11): 1153-1162.
DOI PMID |
| [18] |
Rodrigues t A D, Arruda E J D, Fernandes M F, et al. Copper II-polar amino acid complexes: toxicity to bacteria and larvae of Aedes aegypti[J]. Anais da Academia Brasileira de Ciências, 2017, 89(3): 2273-2280.
DOI URL |
| [19] | Gutiérrez J-C, de Francisco P, Amaro F, et al. Chapter 22-structural and functional diversity of microbial metallothionein genes[M]// Das S, Dash HR.Microbial Diversity in the Genomic Era. America; Academic Press. 2019: 387-407. |
| [20] | Smirnova G V, Oktyabrsky O N. Glutathione in bacteria[J]. Biochemistry, 2005, 70(11): 1199-1211. |
| [21] | Gutiérrez JC, Francisco P, Amaro F, et al. Structural and functional diversity of microbial metallothionein genes[M]. Microbial Diversity in the Genomic Era Academic Press, 2019, 22(1): 387-407. |
| [22] |
Santiago S, E L C, Omar B, et al. Molecular mechanisms for the reaction between -OH radicals and proline: Insights on the role as reactive oxygen species scavenger in plant stress[J]. Journal of Physical Chemistry B, 2014, 118(1): 37-47.
DOI URL |
| [23] |
Takagi H. Proline as a stress protectant in yeast: physiological functions, metabolic regulations, and biotechnological applications[J]. Applied Microbiology Biotechnology, 2008, 81(2): 211-223.
DOI URL |
| [24] |
Freedman J H, Ciriolo M R, Peisach J. The role of glutathione in copper metabolism and toxicity[J]. Journal of Biological Chemistry, 1989, 264(10): 5598-5605.
PMID |
| [25] |
Newton G L, Arnold K, Price M S, et al. Distribution of thiols in microorganisms: mycothiol is a major thiol in most actinomycetes[J]. Journal of Bacteriology, 1996, 178(7): 1990-1995.
PMID |
| [26] | 许剑. 微紫青霉菌铜抗性特征和基于转录组测序的抗铜网络研究[D]. 南宁: 广西大学, 2018. |
| XU Jian. The study on characteristics of copper resistance and anti copper network based on rna-sequencing of Penicillium Janthinellum[D]. Nanning: Guangxi University, 2018. | |
| [27] |
Klein J S, Lewinson O. Bacterial ATP-driven transporters of transition metals: physiological roles, mechanisms of action, and roles in bacterial virulence[J]. Metallomics Integrated Biometal Science, 2011, 3(11): 1098-1108.
DOI URL |
| [28] | Wilkens S. Structure and mechanism of ABC transporters[J]. F1000 Prime Reports, 2015, 7(14): 7-14. |
| [29] |
Peng C, Huang D, Shi Y, et al. Comparative transcriptomic analysis revealed the key pathways responsible for organic sulfur removal by thermophilic bacterium Geobacillus thermo glucosidasius W-2-Science Direct[J]. The Science of the Total Environment, 2019, 676(323): 639-650.
DOI URL |
| [30] |
Benabdellah K, Valderas A n-Aguilar C, et al. GintABC1 encodes a putative ABC transporter of the MRP subfamily induced by Cu, Cd, and oxidative stress in Glomus intraradices[J]. Mycorrhiza, 2010, 20(2): 137-146.
DOI URL |
| [31] |
Ota Irene M, Varshavsky A. A yeast protein similar to bacterial two-component regulators[J]. Science, 1993, 262(5133): 566-569.
PMID |
| [32] |
Caille O, Rossier C, Perron K. A copper-activated two-component system interacts with zinc and imipenem resistance in Pseudomonas aeruginosa[J]. Journal of Bacteriology, 2007, 189: 4561-4568.
DOI PMID |
| [33] | Yamamoto K, Ishihama A. Characterization of copper-inducible promoters regulated by CpxA/CpxR in Escherichia coli[J]. Bioscience, Biotechnology, and Biochemistry, 2006, 70(7): 1688-1695. |
| [34] |
Yamamoto K, Ishihama A. Transcriptional response of Escherichia coli to external copper[J]. Molecular Microbiology, 2005, 56(1): 215-227.
PMID |
| [35] |
Waidner B, Melchers K, St hler F N, et al. The Helicobacter pylori CrdRS two-component regulation system (HP1364/HP1365) is required for copper-mediated induction of the copper resistance determinant CrdA[J]. J Bacteriol, 2005, 187(13): 4683-4688.
PMID |
| [36] |
Zeng Y, Charkowski A O. The Role of ATP-Binding Cassette Transporters in Bacterial Phytopathogenesis[J]. Phytopathology, 2021, 111(4): 600-610.
DOI URL |
| [1] | WEI Wei, SHAN Shouming, XU Wendi, LI Guangzong. Transcriptome analysis of callus at rooting stage in tissue culture of vitis amurensis 'shuangyou' [J]. Xinjiang Agricultural Sciences, 2023, 60(6): 1451-1459. |
| [2] | WANG Yejian, YANG Jie, LIANG Xiaoling, Abulati Abra, HAN Denxu, XI Haojiang, LIU Jun, LI Mingdong. Transcriptome Analysis of Floral Organ Differentiation Stage of Two Maize Inbred Lines under Drought Stress [J]. Xinjiang Agricultural Sciences, 2020, 57(9): 1578-1585. |
| [3] | WANG Zhen-Dong, LU Xiao-Yan, TU Wen-Wen, HE Chen-Chen. Transcriptome Sequencing Analysis of Leaf and Root of Sour Jujube Seedlings under NaCl Stress Alleviated by Exogenous CaCl2 [J]. Xinjiang Agricultural Sciences, 2019, 56(6): 1052-1062. |
| [4] | HUANG Cheng-wen, WANG Xi-dong, LIU Jun, YANG Dong-sheng, Gunisaha Matoheti, Reziwan Aobuli. Effects of Two Types of Expression Vectors on Biological Characteristics of Acidovorax citrulli [J]. Xinjiang Agricultural Sciences, 2018, 55(3): 457-467. |
| [5] | ZHANG Yi-yuan, GUO Yang-hua, WANG Cong-hui, TANG Hong, NAN Hai-yan, WANG Li-min, ZHOU Pin. Induction and Transcriptomics Analysis of Ovine iPS Cells [J]. Xinjiang Agricultural Sciences, 2018, 55(11): 2142-2149. |
| [6] | BAO Qiu-juan, ZHANG Li-li, Hainar Wulazibai, ZHANG Fu-chun. Analysis of DNA Damage Repair Related Genes in Drought Stress Cotton Transcriptome [J]. Xinjiang Agricultural Sciences, 2017, 54(11): 1999-2005. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||