

Xinjiang Agricultural Sciences ›› 2023, Vol. 60 ›› Issue (2): 493-500.DOI: 10.6048/j.issn.1001-4330.2023.02.029
• Forestry·Prataculture·Animal Husbandry Veterinarian • Previous Articles Next Articles
Nasibai Abuduwahapu( ), GAO Xiaojuan, Kunduziayi Abudushalamu, LI Jiangwei(
), GAO Xiaojuan, Kunduziayi Abudushalamu, LI Jiangwei( )
)
Received:2022-06-23
Online:2023-02-20
Published:2023-03-31
Correspondence author:
LI Jiangwei (1967-), male, Xuchang, Henan province. Ph.D., researcher, master superivisor,(E-mail)jwli67@sina.com
Supported by:
娜斯拜·阿卜杜瓦哈普( ), 高晓娟, 坤杜孜阿依·阿布都沙拉木, 李江伟(
), 高晓娟, 坤杜孜阿依·阿布都沙拉木, 李江伟( )
)
通讯作者:
李江伟(1967-),男,新疆乌鲁木齐人,教授,硕士生导师,研究方向为抗体工程及肿瘤免疫治疗,(E-mail)jwli67@sina.com
作者简介:娜斯拜·阿卜杜瓦哈普(1994-),女,新疆奇台人,硕士研究生,研究方向为生化与分子生物学,(E-mail)1535313706@qq.com
基金资助:CLC Number:
Nasibai Abuduwahapu, GAO Xiaojuan, Kunduziayi Abudushalamu, LI Jiangwei. Cloning and Sequence Analysis of cDNA of C3d Gene of Bactrian Camel from Xinjiang[J]. Xinjiang Agricultural Sciences, 2023, 60(2): 493-500.
娜斯拜·阿卜杜瓦哈普, 高晓娟, 坤杜孜阿依·阿布都沙拉木, 李江伟. 新疆双峰驼C3d基因cDNA的克隆与序列分析[J]. 新疆农业科学, 2023, 60(2): 493-500.
Add to citation manager EndNote|Ris|BibTeX
URL: http://www.xjnykx.com/EN/10.6048/j.issn.1001-4330.2023.02.029
| 引物 Primer | 序列(5'-3') Sequence(5'-3') | C3中位置 Location in the C3 | 产物长度 The length of product(bp) |
|---|---|---|---|
| C3d-primer-F | cgatccatggTAAAGCACCTCATCGTGACCCCGT | 3 086 | 909 |
| C3d-primer-R | attactcgagGCGGCTGGGCAGGTGGATGGAC | 3 995 | 909 |
Table 1 The primers of C3 gene of Bactrian camel in Xinjiang were amplified by PCR
| 引物 Primer | 序列(5'-3') Sequence(5'-3') | C3中位置 Location in the C3 | 产物长度 The length of product(bp) |
|---|---|---|---|
| C3d-primer-F | cgatccatggTAAAGCACCTCATCGTGACCCCGT | 3 086 | 909 |
| C3d-primer-R | attactcgagGCGGCTGGGCAGGTGGATGGAC | 3 995 | 909 |
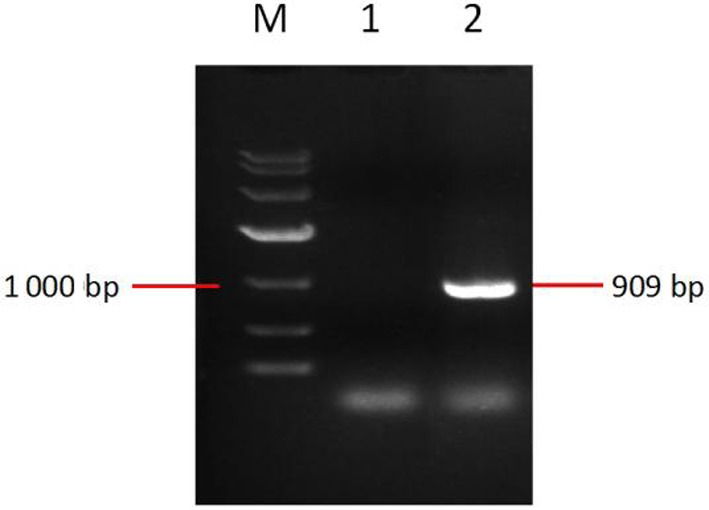
Fig.2 Detection of Bactrian camel C3D gene PCR product by gel electrophoresis Note:M:10000 bp DNA marker; 2:1.Negative control; 3:Bactrian camel C3D gene PCR products
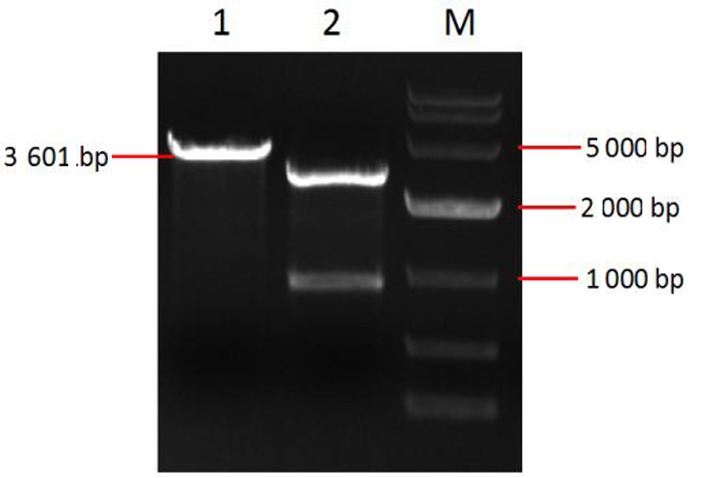
Fig.3 identification of pmd18-t-c3d plasmid by Nco I&Xho I double digestion Note:M:10000 bp DNA marker; 1: Single enzyme digestion products of pMD18-T-C3d; 2: Double enzyme digestion products of pMD18-T-C3d
| 结构分析 Structure analysis | 预测结果 Predicted results |
|---|---|
| α-螺旋 α-Screw | 12-32 37-44 48-54 110-114 122-140 157-162 176-204 217-227 254-268 273-280 |
| β-折叠 β-fold | 1-12 59-93 106-109 163-197 172-175 232-244 276-282 |
| T-转角 T-corner | 33-36 44-47 55-58 96-99 102-105 118-121 141-144 146-149 153-156 168-171 205-208 228-231 246-253 269-272 283-286 |
| 卷曲区 Crimp zone | 1-2 43-45 55-56 58 96-97 99 102 105-106 115-118 145-148 151-153 155-156 168 195 200-201 205 209-210 231-235 240 249-253 268-269 283-284 |
| 亲水区 Hydrophilic zone | 11-32 33-49 58-60 92-104 105-109 113-126 131-144 145-155 157-160 161-162 163-170 171-172 180-181 183-186 187-189 194-216 230-232 234-235 237-254 263-277 282-283 284-287 |
| 柔韧性 Flexibility | 14-18 23-27 33-39 43-45 56-59 79-81 93-99 102-106 115-123 136-139 142-157 167-169 183-186 195-210 215-216 227-234 240-253 266-274 |
| 抗原性 Antigenicity | 12-19 12-37 41-48 55-60 91-107 113-126 135-160 164-172 181-188 189-190 197-213 215-217 227-235 239-254 265-278 283-287 |
| 表面可及性 Surface accessibility | 11-19 23-28 34-39 43-47 93-98 117-125 165-170 197-212 239-245 264-274 |
Table 2 Analysis of secondary structure of complement C3d of Bactrian camel in Xinjiang
| 结构分析 Structure analysis | 预测结果 Predicted results |
|---|---|
| α-螺旋 α-Screw | 12-32 37-44 48-54 110-114 122-140 157-162 176-204 217-227 254-268 273-280 |
| β-折叠 β-fold | 1-12 59-93 106-109 163-197 172-175 232-244 276-282 |
| T-转角 T-corner | 33-36 44-47 55-58 96-99 102-105 118-121 141-144 146-149 153-156 168-171 205-208 228-231 246-253 269-272 283-286 |
| 卷曲区 Crimp zone | 1-2 43-45 55-56 58 96-97 99 102 105-106 115-118 145-148 151-153 155-156 168 195 200-201 205 209-210 231-235 240 249-253 268-269 283-284 |
| 亲水区 Hydrophilic zone | 11-32 33-49 58-60 92-104 105-109 113-126 131-144 145-155 157-160 161-162 163-170 171-172 180-181 183-186 187-189 194-216 230-232 234-235 237-254 263-277 282-283 284-287 |
| 柔韧性 Flexibility | 14-18 23-27 33-39 43-45 56-59 79-81 93-99 102-106 115-123 136-139 142-157 167-169 183-186 195-210 215-216 227-234 240-253 266-274 |
| 抗原性 Antigenicity | 12-19 12-37 41-48 55-60 91-107 113-126 135-160 164-172 181-188 189-190 197-213 215-217 227-235 239-254 265-278 283-287 |
| 表面可及性 Surface accessibility | 11-19 23-28 34-39 43-47 93-98 117-125 165-170 197-212 239-245 264-274 |
| 物种 Species | 分子量 Molecular weight | 等电点 Isoelectric point | pH7时的电荷 The charge at pH7 | 疏水性平均值 Mean hydrophobicity | 抗原性指数 Antigenicity index |
|---|---|---|---|---|---|
| 新疆双峰驼Xinjiang Bactrian camel | 32 185.02 | 6.4 | -3.13 | 0.13 | 0.53 |
| 双峰驼Bactrian camel | 33 951.1 | 6.25 | -2.99 | 0.12 | 0.43 |
| 单峰驼Dromedary camel | 33 951.1 | 6.25 | -2.99 | 0.12 | 0.43 |
| 羊驼Alpaca | 33 865.05 | 6.7 | -0.99 | 0.1 | 0.39 |
| 野驼Wild camel | 33 670.65 | 6.03 | -3.99 | 0.07 | 0.44 |
| 人People | 33 915 | 6.37 | -1.49 | 0.16 | 0.39 |
| 鼠Rat | 34 015.9 | 5.49 | -6.15 | 0.18 | 0.4 |
| 猪Pig | 34 160.23 | 6.13 | -3.15 | 0.19 | 0.47 |
| 家兔Rabbit | 34 255.28 | 7.85 | 1.84 | 0.2 | 0.37 |
| 牛Cattle | 34 457.36 | 7.38 | 0.84 | 0.36 | 0.5 |
Table 3 Analysis of complement C3d protein of Bactrian camel in Xinjiang
| 物种 Species | 分子量 Molecular weight | 等电点 Isoelectric point | pH7时的电荷 The charge at pH7 | 疏水性平均值 Mean hydrophobicity | 抗原性指数 Antigenicity index |
|---|---|---|---|---|---|
| 新疆双峰驼Xinjiang Bactrian camel | 32 185.02 | 6.4 | -3.13 | 0.13 | 0.53 |
| 双峰驼Bactrian camel | 33 951.1 | 6.25 | -2.99 | 0.12 | 0.43 |
| 单峰驼Dromedary camel | 33 951.1 | 6.25 | -2.99 | 0.12 | 0.43 |
| 羊驼Alpaca | 33 865.05 | 6.7 | -0.99 | 0.1 | 0.39 |
| 野驼Wild camel | 33 670.65 | 6.03 | -3.99 | 0.07 | 0.44 |
| 人People | 33 915 | 6.37 | -1.49 | 0.16 | 0.39 |
| 鼠Rat | 34 015.9 | 5.49 | -6.15 | 0.18 | 0.4 |
| 猪Pig | 34 160.23 | 6.13 | -3.15 | 0.19 | 0.47 |
| 家兔Rabbit | 34 255.28 | 7.85 | 1.84 | 0.2 | 0.37 |
| 牛Cattle | 34 457.36 | 7.38 | 0.84 | 0.36 | 0.5 |
| [1] | A A Wahid, R W Dunphy, et al. Insights into the Structure-Function Relationships of Dimeric C3d Fragments[J]. Frontiers in Immunology, 2021(12):1-13.[H1] |
| [2] |
Rasmussen K J, Skjodt M O, Vitved L, et al. A novel antihuman C3d monoclonal antibody with specificity to the C3d complement split product[J]. Journal of Immunological Methods, 2017, 444:51-55.
DOI PMID |
| [3] | 牛明福, 范逸文, 杜梦璇, 等. 鸡γ干扰素与补体C3d主要片段的串联表达及活性测定[J]. 中国免疫学杂志, 2020, 36(6):28-32. |
| NIU Mingfu, FAN Yiwen, DU Mengxuan, et al. Tandem expression and activity test of chicken interferon-γ and complement C3 d major fragments[J]. Chinese Journal of Immunology, 2020, 36(6):28-32. | |
| [4] | Buhlmann D, Eberhardt H U, Medyukhina A, et al. FHR3 Blocks C3d-Mediated Coactivation of Human B Cells[J]. Journal of Immunology, 2016:620-630. |
| [5] | 郑秀惠, 李力, 郭建新, 等. HPV16 E6E7与C3d3 融合基因真核表达质粒的构建及表达[J]. 重庆医学, 2010, 39(1):40-42. |
| ZHENG Xiuhui, LI Li, GUO Jianxin, et al. Construction and expression of eukaryotic expression plasmid of HPV16 E6E7 and C3d3 fusion gene[J]. Chongqing Med, 2010, 39(1):40-42. | |
| [6] | 赵玉斐, 章振华, 李永清, 等. 鸡补体C3d与禽流感病毒M2融合蛋白的原核表达与反应原性的分析[J]. 中国预防兽医学报, 2010: 1-7. |
| ZHAO Yufei, ZHANG Zhenhua, LI Yongqing, et al. Prokaryotic expression and reactivity analysis of fusion protein of Chicken complement C3d and Avian influenza virus M2[J]. Chinese Journal of Preventive Veterinary Medicine, 2010: 1-7. | |
| [7] | 孙盈, 闫敏鑫, 黄秀芬, 等. 表达鸡补体因子C3d基因重组杆状病毒的构建及初步应用[J]. 中国兽医学报, 2015, 35(8):1254-1259. |
| SUN Ying, YAN Minxin, HUANG Xiufen, et al. Construction and preliminary application of recombinant baculovirus expressing chicken complement factor C3d gene[J]. Chinese Journal of Veterinary Medicine, 2015, 35(8):1254-1259. | |
| [8] | De Groot A S, Ross T M, Levitz L, et al. C3d adjuvant effects are mediated through the activation of C3d-specific autoreactive T cells[J]. Immunology & Cell Biology, 2015, 93(2):189-197. |
| [9] | Zhang Y, Guo J, Ning L, et al. The molecular mechanism of pH regulating C3d-CR2 interactions: insights from molecular dynamics simulation[J]. Chemical Biology & Drug Design, 2018:1-11. |
| [10] |
S Muylder ma ns. Applications of Nanobodies[J]. Annual Review of Animal Biosciences, 2021, 9(1):1-10.
DOI URL |
| [11] |
Dempsey P W, Fearon D T. C3d of complement as a molecular adjuvant: bridging innate and acquired immunity[J]. Science, 1996, 271(5247): 348-353.
DOI PMID |
| [12] |
Liu D, Niu Z X. Construction, expression and immunoassay detection of recombinant plasmid encoding fusion protein of Roman chicken complement C3d and Newcastle disease virus F gene[J]. Scandinavian Journal of Immunology, 2010, 68(6):598-606.
DOI URL |
| [13] |
Kulik L, Laskowski J, Renner B, et al. Targeting the Immune Complex-Bound Complement C3d Ligand as a Novel Therapy for Lupus[J]. The Journal of Immunology, 2019, 203(12):1-10.
DOI URL |
| [14] |
Green T D, Newton B R, Rota P A, Xu Y, Robinson H L, Ross T M. C3d enhancement of neutralizing antibodies to measles hemagglutinin[J]. Vaccine, 2002, 20: 242-248.1.
DOI URL |
| [15] |
BrinaS R P, Sundgren A, Sahoo P, et al. Design and synthesis of multifunctional gold nanoparticles bearing tumor-associated glycopeptide antigens as potential cancer vaccines[J]. Bioconjugate Chemistry, 2012, 23(8): 1513-1523.
DOI PMID |
| [16] | 李世军. 牛PLIN1基因转录调控机制及其对前体脂肪细胞增殖、分化和脂类代谢的作用研究[D]. 杨凌: 西北农林科技大学, 2020. |
| LI Shijun. Transcriptional regulation of bovine PLIN1 gene and its effects on proliferation, differentiation and lipid metabolism of precursor adipocytes[D]. Yangling: Northwest A&F University, 2020. | |
| [17] | Elisha R, Verhaar, Andrew W.Woodham, et al. Nanobodies in cancer[J]. Seminars in Immunology, 2020,(52):1-10. |
| [18] | 魏后军, 王芳, 胡波, 等. 兔补体C3d基因cDNA的克隆、鉴定及原核表达[J]. 中国兽医学报, 2013, 33(2):201-205. |
| WEI Houjun, WANG Fang, HU Bo, et al. Cloning, identification and prokaryotic expression of rabbit complement C3d cDNA[J]. Chinese Journal of Veterinary Medicine, 2013, 33(2):201-205. |
| [1] | JIN Juan, Subina Xiaokelaiti, Abudoukayoumu Ayimaiti, YANG Lei, HAO Qing, FAN Dingyu. Cloning and sequence analysis of expansin gene ZjEXPA8 in Huizao [J]. Xinjiang Agricultural Sciences, 2023, 60(9): 2223-2230. |
| [2] | WANG Man, ZHANG Zheng, Yilidana Dilixiati, WU Bin. Cloning and bioinformatics analysis of VvGST1 from Munage tab.grapes [J]. Xinjiang Agricultural Sciences, 2023, 60(8): 1922-1930. |
| [3] | GENG Feifei, MENG Chaomin, QING Guixia, ZHOU Jiamin, ZHANG Fuhou, LIU Fengju. Cloning and expression analysis of phosphorus efficient gene GhMYB4 in Gossypium hirsutum L. [J]. Xinjiang Agricultural Sciences, 2023, 60(6): 1406-1412. |
| [4] | SHANG Jing, PANG Hongbo, WANG Lanlan, LI Xuemei, WANG Yanqiu, LI Yueying. Study on the relationship between auxin and sorghum heterosis [J]. Xinjiang Agricultural Sciences, 2023, 60(4): 841-846. |
| [5] | YI Yuanyang, XIE Fang, TIAN Shiying, ZHANG Zhidong, GU Meiying, PENG Xiaowu. Effect of Organic Fertilizer of Thioerythromycin Residue on Soybean Soil Drug-resistant Bacteria and Their Resistance Genes [J]. Xinjiang Agricultural Sciences, 2023, 60(2): 424-431. |
| [6] | JIN Xiaoye, YE Feng, LIU Liya, MA Xiaojing, LI Xin, SHU Zhan, CHEN Zhuo, ZHONG Qi. Investigation of Grazing Sheep Brucellosis and Implementation of Prevention Measures [J]. Xinjiang Agricultural Sciences, 2023, 60(2): 479-484. |
| [7] | LIU Chenxi, ZHU Yuting, ZHOU Qiang, CHEN Jin, ZHAO Wenjie, ZHENG Kai. Cloning and bioinformatics analysis of β-carotene isomerase GbD27-6 gene in sea island cotton [J]. Xinjiang Agricultural Sciences, 2023, 60(12): 2869-2877. |
| [8] | LI Feng, LAI Ganggang, ZHAO Zhihui, CHEN Xiaofei, ZHU Tiansheng. Identification and 16S rRNA gene diversity analysis of salt cedar(Tamarix chinensis)witches'-broom [J]. Xinjiang Agricultural Sciences, 2023, 60(10): 2551-2557. |
| [9] | WANG Jingyi, ZUO Changgeng, NIU Xinxiang, YANG Hongmei, CHU Min, WANG Ning, LIN Qing, WANG Youwu, LOU Kai, SHI Yingwu. The Quantity and Activity of Biocontrol Bacteria in Cotton Field Soil and the Correlation of Disease Prevention [J]. Xinjiang Agricultural Sciences, 2023, 60(1): 178-184. |
| [10] | LI Min, MA Ying, Huercha , HE Wenwen, SHI Qianyun, Alimujiang Jiapaer, JIANG Qian, Bayinchahan . Expression of Dm86 Gene of Dermacenter Marginatus and Analysis of Immunogenicity [J]. Xinjiang Agricultural Sciences, 2022, 59(9): 2324-2332. |
| [11] | WANG Zhong, FAN Zheru, ZHANG Yueqiang, LI Jianfeng, GAO Xin, SHI Jia. Cloning and Expression Analysis of Chlorophyll a Oxygenase Gene in Wheat [J]. Xinjiang Agricultural Sciences, 2021, 58(9): 1569-1576. |
| [12] | YANG Bo, CHE Yuhong, GUO Chunmiao, Mubareke·Ayoupu, WU Jingrong, DU Juan. Cloning and bioinformation analysis of 4CL gene in Cydonia oblonga Mill [J]. Xinjiang Agricultural Sciences, 2021, 58(6): 1055-1063. |
| [13] | GUAN Lihui, LIU Ping, DANG Wenfang, YANG Hongmei, NIU Xinxiang, CHU Ming, LI Ping, GAO Yan, ZENG Jun, HUO Xiangdong, ZHANG Tao, LIN Qing, Mahemuti Outikuer, LI Yuguo, LOU Kai, SHI Yingwu. Establishment and Spatiotemporal Distribution Characteristics of Rhizosphere Soil Archaea of Cotton by Real-time Fluorescent TaqMan-quantitative PCR [J]. Xinjiang Agricultural Sciences, 2021, 58(3): 450-456. |
| [14] | MIN Kaili, CHAO Xiangbao, TENG Lu, CAI Yongsheng, LEI Huichen, YAN Zhongjian, ZHENG Kai, CHEN Quanjia. Cloning and Expression Analysis of GbHCT10 Gene of Gossypium barbadense L. [J]. Xinjiang Agricultural Sciences, 2021, 58(2): 206-215. |
| [15] | YANG Jieping, ZHOU Li, Ma Li, QUAN Shaowen, Qin Yang, NIU Jianxin. Detection of Pear Virus by High-Throughput Sequencing Technology [J]. Xinjiang Agricultural Sciences, 2020, 57(8): 1503-1513. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||