

新疆农业科学 ›› 2023, Vol. 60 ›› Issue (10): 2396-2403.DOI: 10.6048/j.issn.1001-4330.2023.10.007
玛依拉·玉素音( ), 阳妮, 徐海江, 杨延龙, 张大伟, 李春平, 石必显, 赖成霞(
), 阳妮, 徐海江, 杨延龙, 张大伟, 李春平, 石必显, 赖成霞( )
)
收稿日期:2023-02-10
出版日期:2023-10-20
发布日期:2023-11-01
通信作者:
赖成霞(1972-),女,新疆乌鲁木齐人,副研究员,研究方向为棉花分子育种,(E-mail)作者简介:玛依拉·玉素音(1987-),女,新疆乌鲁木齐人,助理研究员,研究方向为棉花分子育种,(E-mail)119327132@qq.com
基金资助:
Mayila Yusuyin( ), YANG Ni, XU Haijiang, YANG Yanlong, ZHANG Dawei, LI Chunping, SHI Bixian, LAI Chengxia(
), YANG Ni, XU Haijiang, YANG Yanlong, ZHANG Dawei, LI Chunping, SHI Bixian, LAI Chengxia( )
)
Received:2023-02-10
Online:2023-10-20
Published:2023-11-01
Correspondence author:
LAI Chengxia (1972-), female, native place: Urumqi, associate researcher, research direction: cotton molecular breeding, (E-mail)Supported by:摘要:
【目的】 筛选分析受棉铃虫诱导的棉花糖基转移酶(Glycosyltransferase)UGT基因家族,为UGTG01、UGTG02、UGTGo4、UGTGo6、UGTGo7基因功能深入研究奠定基础。【方法】 以受棉铃虫诱导筛选出的棉花糖基转移酶UGT基因组数据库为基础,利用生物信息学方法分析棉花糖基转移酶UGT基因家族的理化性质、系统进化树、蛋白质的二级结构预测以及保守基序与结构域。【结果】 从棉花基因组中搜索到5个推定的UTG基因家族成员,其氨基酸数目介于468~534个,理论等电点分布在4.97~5.58,相对分子质量区域为52 743.1~60 208.62,亚细胞定位在细胞质外,均为亲水性蛋白质。基因结构均含有1个外显子,二级结构主要由α-螺旋、无规则卷曲为主,均含3个Motif基序和有Glycosyltransferase_GTB-type superfamily结构域。5个基因在根、茎、子叶、真叶、花、棉桃中具有明显的组织表达差异性,5个基因与UGT73C家族基因,水稻的UGT707A3、大豆的GmUGT在进化树的同一分支上。【结论】 糖基转移酶基因在不同组织中的表达具有显著差异,5个糖基转移酶与UGT73C、UGT707A3、GmUGT都处于进化树的同一个分支上,其可能具有相似的抗虫功能,能通过调控皂苷元、类黄酮、柚皮素等物质响应棉花的抗虫性。
中图分类号:
玛依拉·玉素音, 阳妮, 徐海江, 杨延龙, 张大伟, 李春平, 石必显, 赖成霞. 受棉铃虫诱导的棉花糖基转移酶UGT基因家族的生物信息学及表达分析[J]. 新疆农业科学, 2023, 60(10): 2396-2403.
Mayila Yusuyin, YANG Ni, XU Haijiang, YANG Yanlong, ZHANG Dawei, LI Chunping, SHI Bixian, LAI Chengxia. Bioinformatics and expression analysis of UGT gene family of cotton glycosyltransferase damaged by Helicoverpa armigera[J]. Xinjiang Agricultural Sciences, 2023, 60(10): 2396-2403.
| 基因Gene | 正向引物Forward primer(5'-3') | 反向引物Reversr primer(5'-3') |
|---|---|---|
| histone 3 | GGTGGTGTGAAGAAGCCTCAT | AATTTCACGAACAAGCCTCTGGAA |
| UGTG01 | TTTCGCTCTGCAATAGGACA | AGCGATCTCCTTTAACTGACC |
| UGTG02 | ATCAAGGGACTAATGCGAAG | CAAATCCAAGCACCTAACGA |
| UGTGo4 | CTCAACCGTTTAACACCTCC | GCCTCTTCCTTTTACCCTT |
| UGTGo6 | ACGCTAAAATCCTTATTAACACC | AGTTAAAGAATGAAGATGGGACT |
| UGTGo7 | GTCATCTAAACTCTTTCCCACT | AGATTCAATAAAGGTCCCACT |
表1 定量PCR 引物序列
Tab.1 Primer sequences for RT-qPCR
| 基因Gene | 正向引物Forward primer(5'-3') | 反向引物Reversr primer(5'-3') |
|---|---|---|
| histone 3 | GGTGGTGTGAAGAAGCCTCAT | AATTTCACGAACAAGCCTCTGGAA |
| UGTG01 | TTTCGCTCTGCAATAGGACA | AGCGATCTCCTTTAACTGACC |
| UGTG02 | ATCAAGGGACTAATGCGAAG | CAAATCCAAGCACCTAACGA |
| UGTGo4 | CTCAACCGTTTAACACCTCC | GCCTCTTCCTTTTACCCTT |
| UGTGo6 | ACGCTAAAATCCTTATTAACACC | AGTTAAAGAATGAAGATGGGACT |
| UGTGo7 | GTCATCTAAACTCTTTCCCACT | AGATTCAATAAAGGTCCCACT |
| 基因 Gene | 蛋白长度 Length(aa) | 分子质量 Molecular weight (kDa) | 等电点 Theoretical pI | 不稳定系数 Instability index | 脂溶性指数 Aliphatic index | 亲水性 Hydrophilic |
|---|---|---|---|---|---|---|
| UGTG01 | 534 | 60 208.62 | 5.22 | 49.66 | 85.77 | -0.183 |
| UGTG02 | 469 | 52 972.26 | 5.01 | 48.29 | 86.84 | -0.202 |
| UGTGo4 | 493 | 54 852.32 | 5.58 | 40.18 | 93.75 | -0.143 |
| UGTGo6 | 469 | 52 660.09 | 5.31 | 40.46 | 84.58 | -0.233 |
| UGTGo7 | 468 | 52 743.1 | 4.97 | 49.85 | 89.12 | -0.166 |
表2 棉花糖基转移酶UGT蛋白基本理化参数
Tab.2 Basic physical and chemical parameters of UGT protein of cotton glycosyltransferase
| 基因 Gene | 蛋白长度 Length(aa) | 分子质量 Molecular weight (kDa) | 等电点 Theoretical pI | 不稳定系数 Instability index | 脂溶性指数 Aliphatic index | 亲水性 Hydrophilic |
|---|---|---|---|---|---|---|
| UGTG01 | 534 | 60 208.62 | 5.22 | 49.66 | 85.77 | -0.183 |
| UGTG02 | 469 | 52 972.26 | 5.01 | 48.29 | 86.84 | -0.202 |
| UGTGo4 | 493 | 54 852.32 | 5.58 | 40.18 | 93.75 | -0.143 |
| UGTGo6 | 469 | 52 660.09 | 5.31 | 40.46 | 84.58 | -0.233 |
| UGTGo7 | 468 | 52 743.1 | 4.97 | 49.85 | 89.12 | -0.166 |
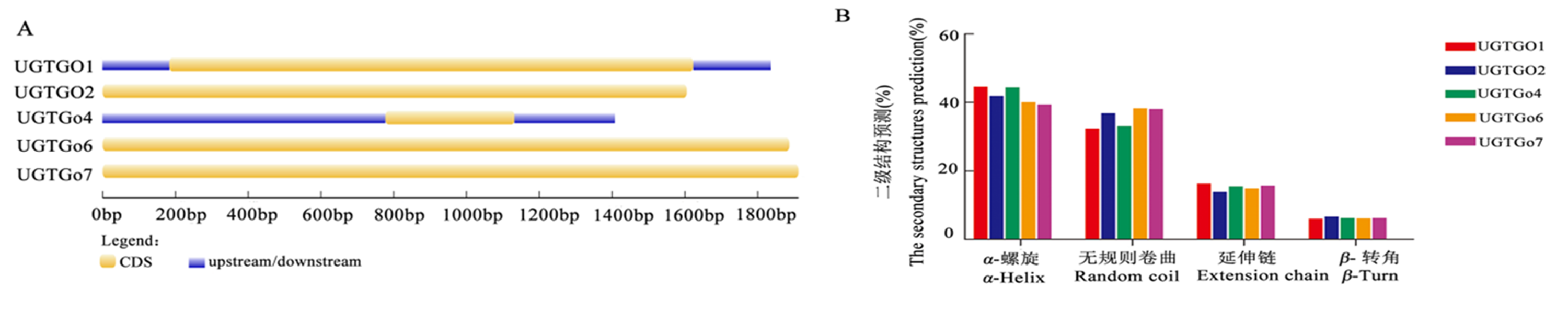
图1 棉花糖基转移酶UGT家族基因结构和二级结构预测 注:A:UGT家族基因结构分析;B: UGT家族基因蛋白质二结构预测
Fig.1 Gene structure analysis and the secondary structures prediction of cotton glycosyltransferase UGT family Note: A:the gene structure analysis of UGT gene family; B: the secondary structures prediction of UGT gene family
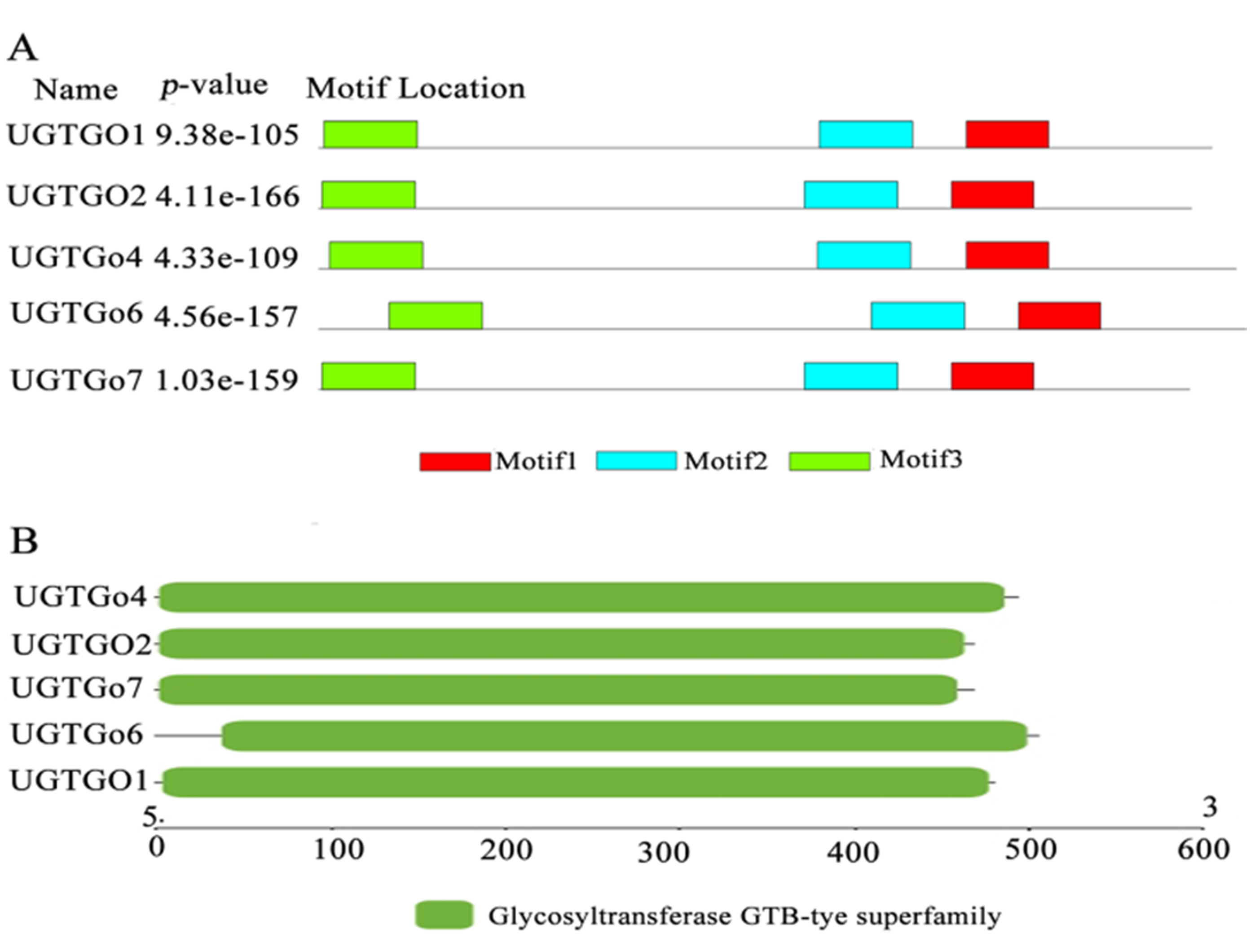
图3 棉花糖基转移酶UGT基因家族蛋白质保守基序和保守结构域 注:A:UGT家族蛋白质保守基序;B:UGT家族基因保守结构域分析
Fig.3 The analysis of protein conserved motifs and domain of cotton glycosyltransferase UGT gene family Note: A:protein conserved motif of UGT gene family;B.Conserved domain analysis of UGT gene family
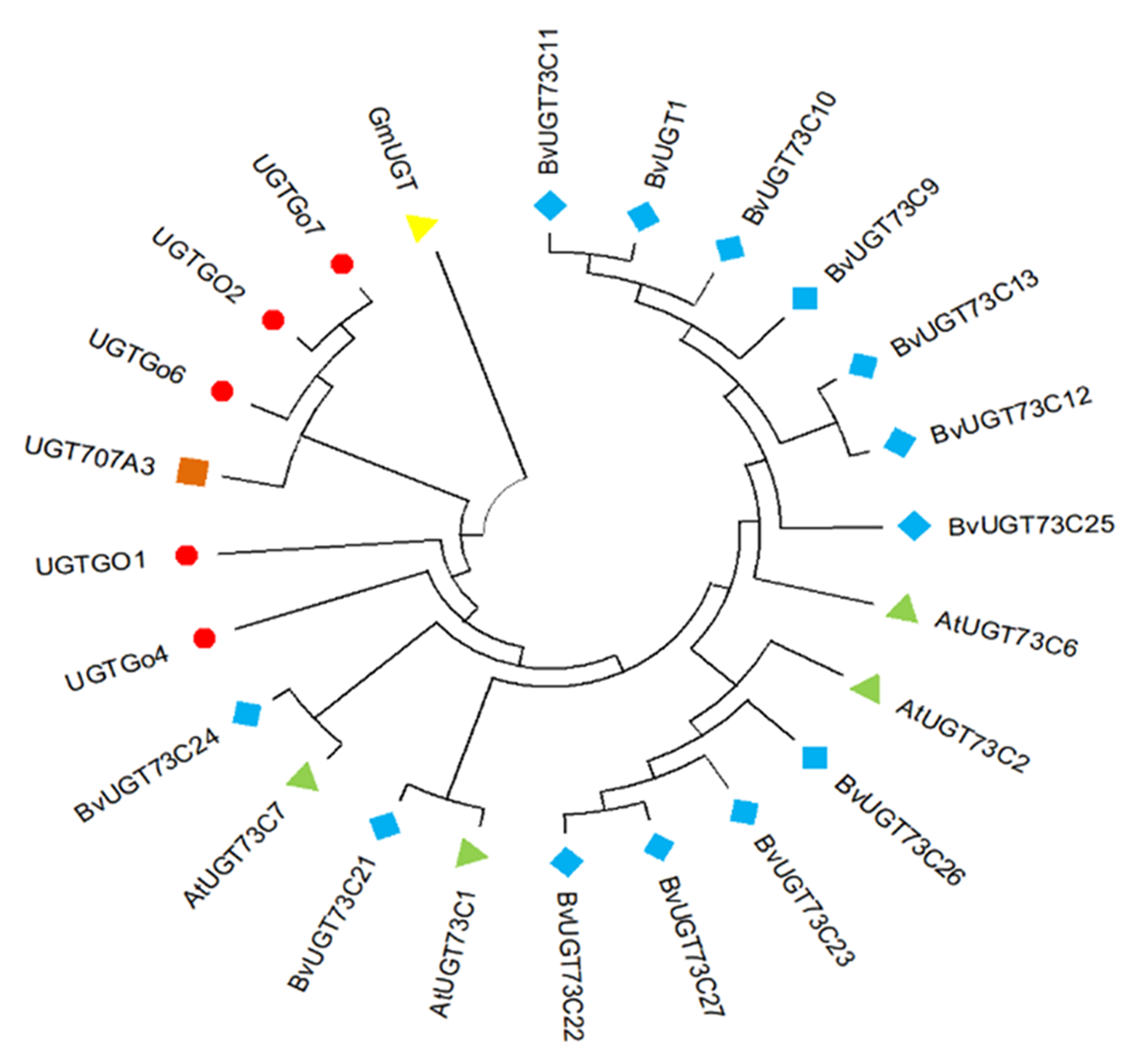
图5 不同植物UGTs分子进化树 注:不同物种UGT不同颜色标记,拟南芥、山芥、水稻、大豆及棉花分别由绿色三角形、蓝色菱形、橙色方形、黄色倒三角形及红色圆形标记。
Fig.5 Phylogenetic tree of different plants UGTs Note:UGTs from the different species were labeled with different colored dots Arabidopsis thaliana, sorrel, rice, soybean, and cotton, represented as green triangles, blue diamonds, orange squares, yellow inverted triangles, and red circles, respectively.
| [1] |
Calvin W, Yang F, Brown SA, et al. Development of Economic Thresholds Toward Bollworm (Lepidoptera: Noctuidae), Management in Bt Cotton, and Assessment of the Benefits From Treating Bt Cotton With Insecticide[J]. Journal of Economic Entomology, 2021, 114(6): 2493-2504.
DOI PMID |
| [2] |
Chen B, Zhang Y, Sun Z, et al. Tissue-specific expression of GhnsLTPs identified via GWAS sophisticatedly coordinates disease and insect resistance by regulating metabolic flux redirection in cotton[J]. The Plant Journal, 2021, 107(3): 831-846.
DOI PMID |
| [3] | 于安东, 刘琳, 龙瑞才, 等. 植物UDP-糖基转移酶(UGT)的功能及应用前景[J]. 植物生理学报, 2022, 58(4): 631-642. |
|
YU Andong, LIU Lin, LONG Ruicai, et al. Function and application prospect of plant UDP-glycosyltransferase (UGT)[J]. Plant Physiology Journal, 2022, 58(4): 631-642.
DOI URL |
|
| [4] |
Rahimi S, Kim J, Mijakovic I, et al. Triterpenoid-biosynthetic UDP-glycosyltransferases from plants[J]. Biotechnology Advances, 2019, 37(7): 107394.
DOI URL |
| [5] |
Rehman H M, Nawaz M A, Shah Z H, et al. Comparative genomic and transcriptomic analyses of Family-1 UDP glycosyltransferase in three Brassica species and Arabidopsis indicates stress-responsive regulation[J]. Scientific reports, 2018, 8(1): 1875.
DOI PMID |
| [6] | Jing T, Zhang N, Gao T, et al. Glucosylation of (Z)-3-hexenol informs intraspecies interactions in plants: A case study in Camellia sinensis[J]. Plant, Cell & Environment. 2019, 42(4): 1352-1367. |
| [7] |
Augustin JM, Drok S, Shinoda T, et al. UDP-glycosyltransferases from the UGT73C subfamily in Barbarea vulgaris catalyze sapogenin 3-O-glucosylation in saponin-mediated insect resistance[J]. Plant Physiology, 160(4): 1881-1895.
DOI URL |
| [8] |
Zhang Y, Guo W, Chen L, et al. CRISPR/Cas9-Mediated Targeted Mutagenesis of GmUGT Enhanced Soybean Resistance Against Leaf-Chewing Insects Through Flavonoids Biosynthesis[J]. Frontiers in Plant Science, 2022, 13:802716.
DOI URL |
| [9] |
Yang Z, Li N, Kitano T, et al. Genetic mapping identifies a rice naringenin O-glucosyltransferase that influences insect resistance[J]. The Plant Journal, 2021, 106(5): 1401-1413.
DOI URL |
| [10] |
Gilbert MK, Bland JM, Shockey JM, et al. A Transcript Profiling Approach Reveals an Abscisic Acid-Specific Glycosyltransferase (UGT73C14) Induced in Developing Fiber of Ligon lintless-2 Mutant of Cotton (Gossypium hirsutum L.)[J]. PLoS ONE, 2013, 8(9): e75268.
DOI URL |
| [11] |
Qin L-X, Rao Y, Li L, et al. Cotton GalT1 Encoding a Putative Glycosyltransferase Is Involved in Regulation of Cell Wall Pectin Biosynthesis during Plant Development. PLoS ONE, 2013, 8(3): e59115.
DOI URL |
| [12] |
Krempl C, Sporer T, Reichelt M, et al. Potential detoxification of gossypol by UDP-glycosyltransferases in the two Heliothine moth species Helicoverpa armigera and Heliothis virescens[J]. Insect Biochemistry and Molecular Biology, 2016, 71:49-57.
DOI PMID |
| [13] |
Kosaka A, Manickavelu A, Kajihara D, et al. Altered gene expression profiles of wheat genotypes against Fusarium head blight[J]. Toxins (Basel), 2015, 7(2): 604-620.
DOI PMID |
| [14] |
Langlois-Meurinne M, Gachon CM, Saindrenan P. Pathogen-responsive expression of glycosyltransferase genes UGT73B3 and UGT73B5 is necessary for resistance to Pseudomonas syringae pv tomato in Arabidopsis[J]. Plant Physiology, 2005, 139(4): 1890-1901.
PMID |
| [15] |
Poppenberger B, Berthiller F, Lucyshyn D, et al. Detoxification of the Fusarium mycotoxin deoxynivalenol by a UDP-glucosyltransferase from Arabidopsis thaliana[J]. Journal of Biological Chemistry, 2003, 278(48): 47905-47914.
DOI PMID |
| [16] | 赖成霞, 玛依拉·玉素音, 高燕, 等. 藏红花酸糖基转移酶UGTCs4基因的克隆、生物信息学及表达分析[J]. 中草药, 2020, 51(4): 1044-1051. |
| LAI Chengxia, Mayila Yusuyin, GAO Yan, et al. Cloning, bioinformatics, expression mode analysis of crocetin glycosyltransferase UGTCs4[J]. Chinese Traditional and Herbal Drugs, 2020, 51(4): 1044-1051. | |
| [17] | 钟晓武, 付强, 邹颉, 等. 烟草糖基转移酶基因NtGT5a的克隆及原核表达. 烟草科技, 2014,(7): 69-74,84. |
| ZHONG Xiaowu, FU Qiang, ZOU Jie, et al. Cloning and Prokaryotic Expression of Glycosyltransferase Gene NtGT5a from Nicotiana tabacum[J]. Tobacco Science and Technology, 2014,(7): 69-74,84. | |
| [18] | 张建军, 文国琴, 刘震, 等. 拟南芥糖基转移酶基因UGT71C5启动子的克隆和功能分析. 四川大学学报(自然科学版), 2011, 48(4): 955-960. |
| ZHANG Jianjun, WEN Guoqin, LIU Zhen, et al. Cloning and functional analysis of UGT71C5 promoterin Arabidopsis thaliana[J]. Journal of Sichuan University (Natural Science Edition), 2011, 48(4): 955-960. | |
| [19] | 张永恩, 王群, 李潮海. 生物信息学在作物抗性研究中的应用展望[J]. 河南农业大学学报, 2005,(4): 25-28. |
| ZHANG Yongen, WANG Qun, LI Haichao. Outlook of the Applications of Bioinformatics in Crop Resistance Research[J]. Journal of Henan Agricultural University, 2005,(4): 25-28. | |
| [20] |
Huang Juan, Pang Chaoyou, Fan Shuli, et al. Genome-wide analysis of the family 1 glycosyltransferases in cotton.[J]. Molecular Genetics and Genomics : MGG, 2015, 290(5): 1805-1818.
DOI PMID |
| [21] | 翟琼, 郑浩霞, 胡博, 等. 花生DNA甲基转移酶基因家族生物信息学分析[J/OL]. 分子植物育种: 1-22[2022-11-19]. |
| ZHAI Qiong, ZHENG Haoxia, HU Bo, et al. Bioinformatics Analysis of Peanut DNA Methyltransferase Gene Family[J]. Molecular Plant Breeding, 1-22[2022-11-19]. | |
| [22] | 李寒, 夏鹏亮, 唐艳红, 等. 小麦OXR基因家族全基因组生物信息学及表达分析[J/OL]. 分子植物育种:1-10[2022-11-19]. |
| LI Han, XIA Pengliang, TANG Yanhong, et al. Genome-wide Bioinformatics and Expression Analysis of OXR Gene Family in Triticum Aestivum[J]. Molecular Plant Breeding, 1-10[2022-11-19]. | |
| [23] | 彭洪娴, 邱洁雅, 惠秋玲, 等. 资阳香橙砧木CjSPL基因家族鉴定及表达模式分析[J/OL]. 生物工程学报:1-19[2022-11-19]. |
| PENG Hongxian, IU Jieya, HUI Qiuling, et al. Identification of CjSPL gene family in Ziyang Xiangcheng rootstock and expression pattern analysis[J]. Chinese Journal of Biotechnology :1-19[2022-11-19]. | |
| [24] |
Wang D, Shi X, Liu D, et al. Transcriptome Profiling Revealed Potentially Critical Roles for Digestion and Defense-Related Genes in Insects' Use of Resistant Host Plants: A Case Study with Sitobion Avenae[J]. Insects, 2020, 11(2): 90.
DOI URL |
| [25] | 李健英. 棉花响应烟粉虱胁迫的分子机制及棉花体细胞胚胎表观基因组学研究[D]. 武汉: 华中农业大学, 2019. |
| LI Jianying. Molecular mechanism of cotton in response to whitefly ifestation and epigenomics basis of cotton somatic embryogenesis[D]. Wuhan: Huazhong Agricultural University, 2019. | |
| [26] | 陈云霄. 玉米和大刍草抗虫性评价及转录组分析[D]. 临汾: 山西师范大学, 2019. |
| CHEN Xiaoyun. Evaluation of Insect Resistance and Transcriptome Analysis of Maize and Teosinte[D]. Linfen: Journal of Shanxi Normal University, 2019. | |
| [27] | 刘龙博, 郑树轩, 郑洁. 石榴低温响应因子CBF基因家族鉴定及其表达分析[J]. 西北农业学报, 2022, 31(9): 1154-1167. |
| LIU Longbo, ZHENG Shuxuan, ZHENG Jie. Identification and Expression Analysis of CBF(C-repeat Binding Factor)Gene Family in Pomegranate(Punica granatum L.)[J]. Acta Agriculturae Boreali-Occidentalis Sinica, 2022, 31(9): 1154-1167. |
| [1] | 邵盘霞, 赵准, 邵武奎, 郝晓燕, 高升旗, 李建平, 胡文冉, 黄全生. 玉米ZmCDPK22基因在干旱胁迫下的表达分析[J]. 新疆农业科学, 2023, 60(6): 1372-1378. |
| [2] | 刘晨曦, 朱雨婷, 周强, 陈瑾, 赵文杰, 郑凯. 海岛棉β-胡萝卜素异构酶GbD27-6基因克隆及表达分析[J]. 新疆农业科学, 2023, 60(12): 2869-2877. |
| [3] | 杨茹薇, 刘易, 李江涛, 古丽米拉·热合木土拉, 罗正乾, 徐琳黎, 沈洪飞. 马铃薯病毒病病原RT-PCR分子鉴定[J]. 新疆农业科学, 2023, 60(12): 3032-3040. |
| [4] | 胡文冉, 赵准, 邵武奎, 黄全生. 陆地棉TCP家族基因鉴定及组织表达分析[J]. 新疆农业科学, 2023, 60(11): 2627-2637. |
| [5] | 才羿, 戴冬洋, 谭海, 王迪, 杨芬, 王岭, 盛云燕. 甜瓜AMS基因结构分析及基因编辑载体构建[J]. 新疆农业科学, 2023, 60(1): 61-68. |
| [6] | 李敏, 马英, 呼尔查, 何文文, 史倩云, 阿力木江·加帕尔, 蒋倩, 巴音查汗. 边缘革蜱Dm 86基因表达及免疫原性分析[J]. 新疆农业科学, 2022, 59(9): 2324-2332. |
| [7] | 戴麒, 徐瑞强, 李宁, 王柏柯, 王娟, 黄少勇, 帕提古丽·艾斯木托拉, 余庆辉. 潘那利番茄GRF基因家族全基因组鉴定分析[J]. 新疆农业科学, 2022, 59(10): 2456-2465. |
| [8] | 孙晓军, 周婷婷, 玉山江·麦麦提, 何伟, 罗文芳, 许建军, 阿布都赛买提·吐尔送. 设施番茄病毒病病原鉴定[J]. 新疆农业科学, 2021, 58(1): 99-106. |
| [9] | 王德钢, 马光皇, 贺鹏鹏, 熊仁次, 曹玉, 张萍, 韩旭, 杨明禄. 香梨优斑螟脂肪酸Δ9去饱和酶基因序列及不同虫态的表达分析[J]. 新疆农业科学, 2020, 57(5): 877-887. |
| [10] | 杨雪梅, 范雪, 田可川, 黄锡霞, 朱桦, 阿布来提·苏来曼, 陈华峰, 何军敏, 赵冰茹, 徐新明, 付雪峰, 田月珍, 吴伟伟, 哈尼克孜·吐拉甫, 杜建文. 苏博美利奴羊胚胎期WNT2基因的组织表达及其生物信息学预测分析[J]. 新疆农业科学, 2019, 56(12): 2320-2328. |
| [11] | 徐航, 全绍文, 马丽, 周丽, 牛建新. 库尔勒香梨PsiERF基因的克隆和表达分析[J]. 新疆农业科学, 2018, 55(9): 1616-1625. |
| [12] | 郝晓燕, 李建平, 常晓春, 足木热木, 高升旗, 陈果, 黄全生. 盐地碱蓬PsaH基因的克隆及生物信息学分析[J]. 新疆农业科学, 2018, 55(7): 1209-1217. |
| [13] | 王玉凯, 赵欢, 牛建新. 库尔勒香梨kfpSPL及其启动子克隆分析[J]. 新疆农业科学, 2018, 55(6): 1017-1026. |
| [14] | 冯亚宁;姜志强;马晓玲;李江伟. 新疆双峰驼纳米抗体噬菌体展示文库多样性的生物信息学分析[J]. , 2016, 53(8): 1533-1538. |
| [15] | 张经博;李波;杨洋;胡文冉;陈方圆;谢丽霞;范玲. 陆地棉CAD基因家族的进化和表达分析[J]. , 2016, 53(7): 1177-1187. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||