

新疆农业科学 ›› 2023, Vol. 60 ›› Issue (11): 2806-2815.DOI: 10.6048/j.issn.1001-4330.2023.11.024
收稿日期:2023-01-30
出版日期:2023-11-20
发布日期:2023-12-07
通信作者:
张利莉(1963-),女,新疆人,教授,博士,硕士生/博士生导师,研究方向为微生物资源与利用,(E-mail)zhanglily@taru.edu.cn
作者简介:张萍(1994-),女,河南人,硕士研究生,研究方向为放线菌资源及其次级代谢产物,(E-mail)zhang_ping16@sina.com
基金资助:
ZHANG Ping( ), LUO Xiaoxia, WAN Chuanxing, ZHANG Lili(
), LUO Xiaoxia, WAN Chuanxing, ZHANG Lili( )
)
Received:2023-01-30
Online:2023-11-20
Published:2023-12-07
Correspondence author:
ZHANG Lili (1963-), female, Xinjiang, professor, research field: microbial resources and utilization,(E-mail)zhanglily@taru.edu.cn
Supported by:摘要:
【目的】 评估菌株TRM SA0054的代谢潜力,解析其基因组信息研究该菌株次级代谢产物基因资源。【方法】 采用Illumina Hiseq测序平台对菌株TRM SA0054进行全基因组测序,并进行生物信息学分析,采用现代分离纯化方法检测鉴定其代谢产物。【结果】 菌株TRM SA0054基因组大小为7.2 Mb,共有6 410个蛋白编码基因,GC含量为73.16%;预测到26个次级代谢基因簇,其中Region 36.1基因簇与Tubercidin(杀结核菌素)基因簇相似性为81%,菌株TRM SA0054发酵能够产生Tubercidin。【结论】 基于基因组挖掘和LC-MS检测验证菌株TRM SA0054具有产生Tubercidin的能力。
中图分类号:
张萍, 罗晓霞, 万传星, 张利莉. 链霉菌TRM SA0054 Tubercidin合成基因簇挖掘[J]. 新疆农业科学, 2023, 60(11): 2806-2815.
ZHANG Ping, LUO Xiaoxia, WAN Chuanxing, ZHANG Lili. Excavating of synthetic gene cluster of Streptomyces TRM SA0054 Tubercidin[J]. Xinjiang Agricultural Sciences, 2023, 60(11): 2806-2815.
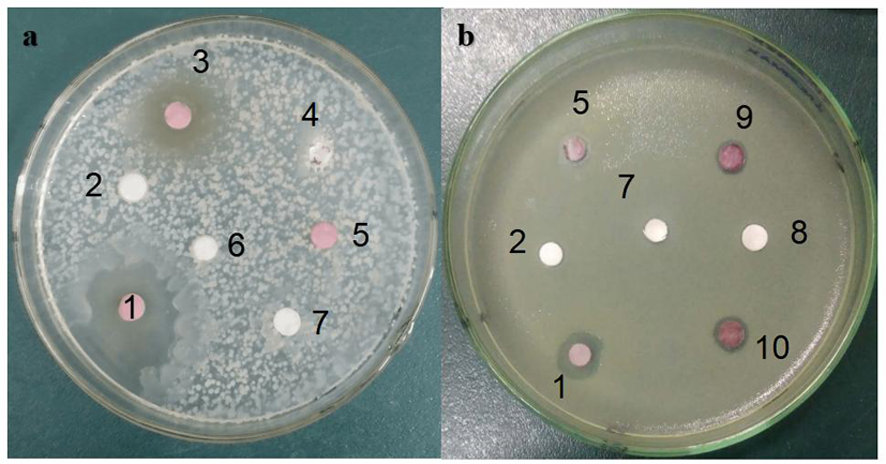
图1 菌株TRM SA0054对靶标病原菌的活性检测 注:a.粪肠球菌(E.faecalis)G+;b.金黄色葡萄球菌(S. aureus)G+;1.甲醇萃取粗提物;2.甲醇空白对照;3.水相粗提物;4.菌体水相;5.菌体甲醇萃取相;6.水空白对照;7.Tubercidin标品(1 mg/mL);8.乙酸乙酯空白对照;9.菌体乙酸乙酯萃取相;10.乙酸乙酯萃取粗提物
Fig. 1 Activity detection of strain TRM SA0054 against target pathogens Note: a.E.faecalis; b.S. aureus;1.Crude extract extracted by methanol; 2.methanol blank control; 3. Aqueous crude extract; 4. Water phase of thallus;5. Methanol extractionof bacteria body; 6.water blank control; 7.Tubercidin standard (1 mg/mL); 8.Ethyl acetate blank control; 9.bacterial body ethyl acetate extraction phase;10.Ethyl acetate extraction of crude extract
| 菌株Strain | S.carminius TRM SA0054 |
|---|---|
| 基因组总长度 Total length of genome (bp) | 7 200 902 |
| CDS基因数 CDS gene number | 6 410 |
| CDS平均长度 Average CDS length (bp) | 954 |
| G+C%含量 G+C% content (%) | 73.15 |
| N50 contig大小 N50 contig size (bp) | 300 464 |
| tRNA数目 tRNA number | 65 |
| rRNA数目 rRNA number | 15 |
表1 S.carminius TRM SA0054 的基因组信息
Tab.1 Genomic information for S.carminius TRM SA0054
| 菌株Strain | S.carminius TRM SA0054 |
|---|---|
| 基因组总长度 Total length of genome (bp) | 7 200 902 |
| CDS基因数 CDS gene number | 6 410 |
| CDS平均长度 Average CDS length (bp) | 954 |
| G+C%含量 G+C% content (%) | 73.15 |
| N50 contig大小 N50 contig size (bp) | 300 464 |
| tRNA数目 tRNA number | 65 |
| rRNA数目 rRNA number | 15 |
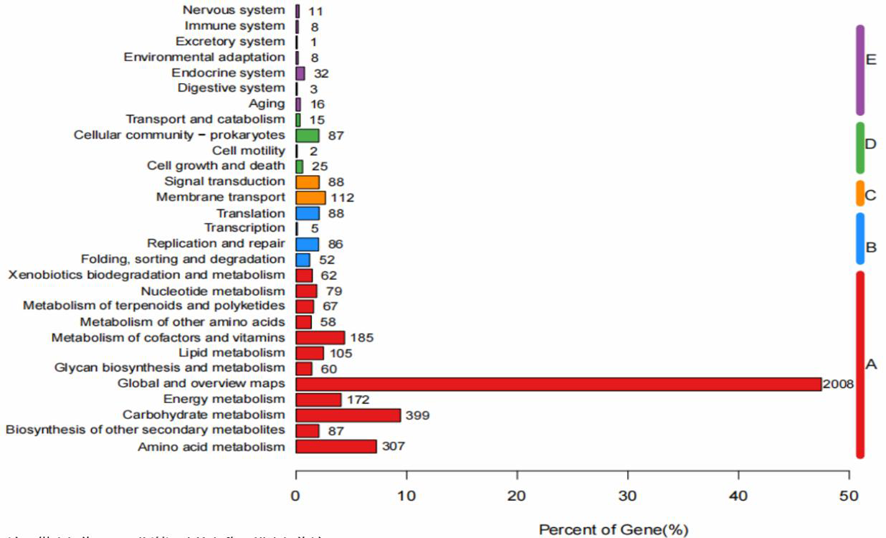
图4 S.carminiusTRM SA0054基因组的KEGG代谢通路分类 注:纵坐标为KEGG代谢通路的名称,横坐标为注释到该通路下的基因个数及其个数占被注释上的基因总数的比例
Fig.4 KEGG metabolic pathway classification of S.carminius TRM SA0054 genome Note: the ordinate is the name of KEGG metabolic pathway, and the ordinate is the number of genes annotated to this pathway and their proportion to the total number of genes annotated
| 基因簇编号 Cluster | 区域 Region | 位置 Location (nt) | 最相似的已知基因簇 Most similar known cluster | 类型 Type | 相似性 Similarity (%) |
|---|---|---|---|---|---|
| 1-407 | 6.1 | 1-21641 | branched-chain fatty acids(支链脂肪酸) | other | 100 |
| 2-419 | 15.1 | 66732-77136 | Ectoine(四氢嘧啶) | other | 100 |
| 3-475 | 41.1 | 25250-66338 | Alkylresorcinol (烷基间苯二酚) | Polyketide | 100 |
| 4-513 | 65.2 | 220132-242865 | Keywimysin (键维霉素) | RiPP | 100 |
| 5-513 | 65.4 | 637199-659916 | SapB | RiPP:Lanthipeptide | 100 |
| 6-502 | 54.1 | 73725-120591 | Undecylprodigiosin (十一烷基灵菌红素) | NRP+ Polyketide | 95 |
| 7-403 | 4.1 | 218512-244303 | Isorenieratene (异戊二烯) | Terpene | 85 |
| 8-462 | 34.1 | 256788-282579 | Isorenieratene (异戊二烯) | Terpene | 85 |
| 9-464 | 36.1 | 13034-33396 | Tubercidin (杀结核菌素) | Other | 81 |
| 10-510 | 62.2 | 204780-240191 | Lagmysin (拉米霉素) | RiPP | 80 |
| 11-403 | 4.2 | 248350-300464 | spore pigmen (孢子色素) | Polyketide | 58 |
| 12-462 | 34.2 | 286626-335712 | spore pigmen (孢子色素) | Polyketide | 58 |
| 13-504 | 56.1 | 107039-149217 | Diazepinomicin | Terpene | 52 |
| 14-513 | 65.3 | 262275-306630 | Melanin (黑色素) | Other | 40 |
| 15-408 | 7.1 | 73518-95460 | Hopene (藿烯) | Terpene | 30 |
| 16-430 | 19.1 | 1-23254 | Microansamycin (微安霉素) | Polyketide | 25 |
| 17-513 | 65.1 | 175916-206600 | Mycosubtilin (抗霉枯草菌素) | NRP + Polyketide | 20 |
| 18-505 | 57.1 | 102245-124388 | lankacidin C (兰卡霉素) | NRP+Polyketide | 20 |
| 19-499 | 51.1 | 1-3282 | Epepothilone B (埃博霉素B) | NRP+Polyketide | 18 |
| 20-510 | 62.1 | 123308-135233 | lipopeptide 8D1-1 / lipopeptide 8D1-2 (脂肽) | NRP | 6 |
| 21-500 | 52.1 | 67269-81582 | Ficellomycin (纤维霉素) | NRP | 5 |
| 22-463 | 35.1 | 63166-83705 | Kosinostatin(越野他汀) | NRP + Polyketide | 4 |
| 23-399 | 1.1 | 69363-81186 | ficellomycin(纤维霉素) | NRP | 3 |
| 24-492 | 48.1 | 69285-89796 | 7-deoxypactamycin (脱氧帕他霉素) | Polyketide | 3 |
| 25-515 | 67.1 | 1-17298 | - | lanthipeptide-class-i | - |
| 26-515 | 67.2 | 183303-194151 | - | RiPP-like | - |
表2 antiSMASH预测S.carminius TRM SA0054的相似度由大到小的基因簇
Tab.2 Gene clusters from large to small similarity predicted by antiSMASH for S.carminius TRM SA0054
| 基因簇编号 Cluster | 区域 Region | 位置 Location (nt) | 最相似的已知基因簇 Most similar known cluster | 类型 Type | 相似性 Similarity (%) |
|---|---|---|---|---|---|
| 1-407 | 6.1 | 1-21641 | branched-chain fatty acids(支链脂肪酸) | other | 100 |
| 2-419 | 15.1 | 66732-77136 | Ectoine(四氢嘧啶) | other | 100 |
| 3-475 | 41.1 | 25250-66338 | Alkylresorcinol (烷基间苯二酚) | Polyketide | 100 |
| 4-513 | 65.2 | 220132-242865 | Keywimysin (键维霉素) | RiPP | 100 |
| 5-513 | 65.4 | 637199-659916 | SapB | RiPP:Lanthipeptide | 100 |
| 6-502 | 54.1 | 73725-120591 | Undecylprodigiosin (十一烷基灵菌红素) | NRP+ Polyketide | 95 |
| 7-403 | 4.1 | 218512-244303 | Isorenieratene (异戊二烯) | Terpene | 85 |
| 8-462 | 34.1 | 256788-282579 | Isorenieratene (异戊二烯) | Terpene | 85 |
| 9-464 | 36.1 | 13034-33396 | Tubercidin (杀结核菌素) | Other | 81 |
| 10-510 | 62.2 | 204780-240191 | Lagmysin (拉米霉素) | RiPP | 80 |
| 11-403 | 4.2 | 248350-300464 | spore pigmen (孢子色素) | Polyketide | 58 |
| 12-462 | 34.2 | 286626-335712 | spore pigmen (孢子色素) | Polyketide | 58 |
| 13-504 | 56.1 | 107039-149217 | Diazepinomicin | Terpene | 52 |
| 14-513 | 65.3 | 262275-306630 | Melanin (黑色素) | Other | 40 |
| 15-408 | 7.1 | 73518-95460 | Hopene (藿烯) | Terpene | 30 |
| 16-430 | 19.1 | 1-23254 | Microansamycin (微安霉素) | Polyketide | 25 |
| 17-513 | 65.1 | 175916-206600 | Mycosubtilin (抗霉枯草菌素) | NRP + Polyketide | 20 |
| 18-505 | 57.1 | 102245-124388 | lankacidin C (兰卡霉素) | NRP+Polyketide | 20 |
| 19-499 | 51.1 | 1-3282 | Epepothilone B (埃博霉素B) | NRP+Polyketide | 18 |
| 20-510 | 62.1 | 123308-135233 | lipopeptide 8D1-1 / lipopeptide 8D1-2 (脂肽) | NRP | 6 |
| 21-500 | 52.1 | 67269-81582 | Ficellomycin (纤维霉素) | NRP | 5 |
| 22-463 | 35.1 | 63166-83705 | Kosinostatin(越野他汀) | NRP + Polyketide | 4 |
| 23-399 | 1.1 | 69363-81186 | ficellomycin(纤维霉素) | NRP | 3 |
| 24-492 | 48.1 | 69285-89796 | 7-deoxypactamycin (脱氧帕他霉素) | Polyketide | 3 |
| 25-515 | 67.1 | 1-17298 | - | lanthipeptide-class-i | - |
| 26-515 | 67.2 | 183303-194151 | - | RiPP-like | - |
| 位点 Locus tag | 氨基酸 Amino acids | 功能预测 Proposed function | 一致性 Identify(%) | 登录号 Accession No. |
|---|---|---|---|---|
| 1-ctg36_17 | 319 | glucosyl-3-phosphoglycerate (葡萄糖基-3-磷酸甘油酸酯) | 92.16 | WP_101256758.1 |
| 2-ctg36_18 | 489 | trehalose-6-phosphate synthase (6-磷酸海藻糖合酶) | 86.71 | WP_093658899.1 |
| 3-ctg36_19 | 290 | trehalose-phosphatase(海藻糖磷酸酶) | 91.29 | WP_031509476.1 |
| 4-ctg36_20 | 240 | hypothetical protein | 62.18 | WP_031509478.1 |
| 5-ctg36_21 | 755 | hypothetical protein | 69.15 | WP_155070928.1 |
| 6-ctg36_22 | 170 | hypothetical protein | 69.41 | WP_1155070929.1 |
| 7-ctg36_23 | 86 | DUF3263 domain-containing protein (DUF3263结构域的蛋白 ) | 82.67 | WP_030218603.1 |
| 8-ctg36_24 | 120 | 6carboxytetrahydropterin synthase (6-羧基四氢蝶呤合酶) | 74.17 | AVW83009.1 |
| 9-ctg36_25 | 181 | 7carboxy7deazaguanine synthase QueE/radical SAM family protein (7-羧基-7-脱氮鸟嘌呤合酶 QueE/ 自由基 SAM 家族蛋白) | 72.66 | AVW83010.1 |
| 10-ctg36_26 | 199 | GTP cyclohydrolase I (GCHI)(GTP环水解酶ⅰ) | 78.28 | AVW83011.1 |
| 11-ctg36_27 | 385 | IMP dehydrogenase ( IMP 脱氢酶) | 78.59 | WP_159742672.1 |
| 12-ctg36_28 | 195 | Phosphoribosylpyrophosphate transferase(磷酸核糖转移酶) | 68.51 | WP_159742671.1 |
| 13-ctg36_29 | 422 | UbiD family decarboxylase (UbiD 家族脱羧酶 ) | 78.01 | WP_159742670.1 |
| 14-ctg36_30 | 168 | NUDIX domaincontaining protein (NUDIX结构域的蛋白质) 水解酶 | 79.76 | WP_159742669.1 |
| 15-ctg36_31 | 327 | Carbohydrate kinase family protein (碳水化合物激酶家族蛋白) | 51.95 | WP_159742668.1 |
| 16-ctg36_32 | 429 | MFS (Major facilitator superfamily) transporter (主要促进者超家族转运蛋白) | 68.39 | WP_159742667.1 |
| 17-ctg36_33 | 249 | HAD family hydrolase | 67.52 | WP_057574898.1 |
| 18-ctg36_34 | 395 | hypothetical protein | 46.22 | WP_155070927.1 |
表3 S.carminiusTRM SA0054基因组中潜在的Tubercidin生物合成基因簇中的功能预测
Tab.3 Function prediction in the potential Tubercidin biosynthesis gene cluster in the S.carminius TRM SA0054 genome
| 位点 Locus tag | 氨基酸 Amino acids | 功能预测 Proposed function | 一致性 Identify(%) | 登录号 Accession No. |
|---|---|---|---|---|
| 1-ctg36_17 | 319 | glucosyl-3-phosphoglycerate (葡萄糖基-3-磷酸甘油酸酯) | 92.16 | WP_101256758.1 |
| 2-ctg36_18 | 489 | trehalose-6-phosphate synthase (6-磷酸海藻糖合酶) | 86.71 | WP_093658899.1 |
| 3-ctg36_19 | 290 | trehalose-phosphatase(海藻糖磷酸酶) | 91.29 | WP_031509476.1 |
| 4-ctg36_20 | 240 | hypothetical protein | 62.18 | WP_031509478.1 |
| 5-ctg36_21 | 755 | hypothetical protein | 69.15 | WP_155070928.1 |
| 6-ctg36_22 | 170 | hypothetical protein | 69.41 | WP_1155070929.1 |
| 7-ctg36_23 | 86 | DUF3263 domain-containing protein (DUF3263结构域的蛋白 ) | 82.67 | WP_030218603.1 |
| 8-ctg36_24 | 120 | 6carboxytetrahydropterin synthase (6-羧基四氢蝶呤合酶) | 74.17 | AVW83009.1 |
| 9-ctg36_25 | 181 | 7carboxy7deazaguanine synthase QueE/radical SAM family protein (7-羧基-7-脱氮鸟嘌呤合酶 QueE/ 自由基 SAM 家族蛋白) | 72.66 | AVW83010.1 |
| 10-ctg36_26 | 199 | GTP cyclohydrolase I (GCHI)(GTP环水解酶ⅰ) | 78.28 | AVW83011.1 |
| 11-ctg36_27 | 385 | IMP dehydrogenase ( IMP 脱氢酶) | 78.59 | WP_159742672.1 |
| 12-ctg36_28 | 195 | Phosphoribosylpyrophosphate transferase(磷酸核糖转移酶) | 68.51 | WP_159742671.1 |
| 13-ctg36_29 | 422 | UbiD family decarboxylase (UbiD 家族脱羧酶 ) | 78.01 | WP_159742670.1 |
| 14-ctg36_30 | 168 | NUDIX domaincontaining protein (NUDIX结构域的蛋白质) 水解酶 | 79.76 | WP_159742669.1 |
| 15-ctg36_31 | 327 | Carbohydrate kinase family protein (碳水化合物激酶家族蛋白) | 51.95 | WP_159742668.1 |
| 16-ctg36_32 | 429 | MFS (Major facilitator superfamily) transporter (主要促进者超家族转运蛋白) | 68.39 | WP_159742667.1 |
| 17-ctg36_33 | 249 | HAD family hydrolase | 67.52 | WP_057574898.1 |
| 18-ctg36_34 | 395 | hypothetical protein | 46.22 | WP_155070927.1 |

图5 Tubercidin生物合成基因簇以及酶促步骤 注:a.Streptomyces tubercidicus NBRC13090的Tubercidin的基因簇结构图;b.Streptomyces carminius TRM SA0054的Tubercidin的基因簇结构图;c.Tubercidin的生物合成途径[20]
Fig.5 Tubercidin biosynthetic gene cluster and enzymatic steps Note: a.Structure diagram of the gene cluster for Tubercidinfrom S.tubercidicus NBRC 13090; b.Structure diagram of the gene cluster for Tubercidin fromS.carminius TRM SA0054; c.Proposed pathway for tubercidin bbiosynthesis[20]
| [1] | Pablo-Gomez-Escribano Juan, Silke Alt, Mervyn J-Bibb. Next Generation Sequencing of Actinobacteria for the Discovery of Novel Natural Products[J]. Marine Drugs, 2016, 14(4). |
| [2] | Arul J P, Bhavanath J. New Dimensions of Research on Actinomycetes: Quest for Next Generation Antibiotics[J]. Frontiers in Microbiology, 2016, 7. |
| [3] | Constanze Paulus, Rebets Yuriy, Tokovenko Bogdan, et al. New natural products identified by combined genomics-metabolomics profiling of marine Streptomyces sp. MP131-18[J]. Scientific reports, 2017, 7(1). |
| [4] |
Hu D, Gao C, Sun C, et al. Genome-guided and mass spectrometry investigation of natural products produced by a potential new actinobacterial strain isolated from a mangrove ecosystem in Futian, Shenzhen, China[J]. Scientific Reports, 2019, 9(1): 823.
DOI PMID |
| [5] | 申国文. Tubercidin通过促进核斑形成增加缺血缺氧状态下心肌凋亡毒性的研究[D]. 桂林: 桂林医学院, 2021. |
| SHEN Guowen. Tubercidin increased the toxicity of myocardial apoptosis under ischemia and hypoxia by promoting nuclear plaque formation[D]. Guilin: Guilin Medical College, 2021. | |
| [6] | 王珊珊, 郭文强, 何宁, 等. 基因组信息指导下茂源链霉菌代谢产物研究[J]. 中国抗生素杂志, 2018, 43(1): 28-34. |
| WANG Shanshan, GUO Wenqiang, HE Ning, et al. Genome guided investigation of metabolites of the strain Streptomyces mobaraensis[J]. Chinese Journal of Antibiotics, 2018, 43 (1): 28-34. | |
| [7] |
Kum E, Nce E. Genome-guided investigation of secondary metabolites produced by a potential new strain Streptomyces BA2 isolated from an endemic plant rhizosphere in Turkey[J]. Arch Microbiol, 2021, 203(5): 2431-2438.
DOI PMID |
| [8] |
Hashimoto T, Hashimoto J, Kagaya N, et al. A novel oxazole-containing tetraene compound, JBIR-159, produced by heterologous expression of the cryptic trans-AT type polyketide synthase biosynthetic gene cluster[J]. The Journal of Antibiotics, (Tokyo), 2021, 74(5): 354-358.
DOI |
| [9] | Wang Y, Xia Z, Liu Z, et al. Streptomyces carminius sp. nov. a novel actinomycete isolated from Sophora alopecuroides in Xinjiang, China[J]. Antonie Van Leeuwenhoek, 2018.DOI:10.1007/s10482-018-1069-x. |
| [10] | Li R Q, Li Y R, Kristiansen Karsten, et al. SOAP: short oligonucleotide alignment program[J]. Bioinformatics (Oxford, England), 2018, 24(5). |
| [11] |
J Besemer, Lomsadze A, Borodovsky M. GeneMarkS: a self-training method for prediction of gene starts in microbial genomes. Implications for finding sequence motifs in regulatory regions[J]. Nucleic Acids Research, 2001, 29(12): 2607-2618.
DOI PMID |
| [12] |
Delcher A L, Bratke K A, Powers E C, et al. Identifying bacterial genes and endosymbiont DNA with Glimmer[J]. Bioinformatics, 2007, 23(6): 673-679.
DOI PMID |
| [13] |
Karin L, Peter H, R dland Einar Andreas, et al. RNAmmer: consistent and rapid annotation of ribosomal RNA genes[J]. Nucleic Acids Research, 2007, 35(9):3100.DOI:10.1093/nar/gkm160.
PMID |
| [14] | Chan P P, Lowe T M. tRNAscan-SE: Searching for tRNA genes in genomic sequences[J]. Other, 2019, 1962.DOI:10.1007/978-1-4939-9173-0_1. |
| [15] | Galperin M Y, Kristensen D M, Makarova K S, et al. Microbial genome analysis: the COG approach[M]. Oxford University Press (OUP), 2019,(4).DOI:10.1093/bib/bbx117. |
| [16] | Ana C, Stefan G, García-Gómez Juan Miguel, et al. Blast2GO: a universal tool for annotation, visualization and analysis in functional genomics research[J]. Bioinformatics, 2005,(18): 18.DOI:10.1093/bioinformatics/bti610. |
| [17] | Kanehisa M, Sato Y, Kawashima M. KEGG mapping tools for uncovering hidden features in biological data[J]. Protein Science, 2021.DOI:10.1002/pro.4172. |
| [18] | Blin K, Shaw S, Kloosterman A M, et al. antiSMASH 6.0: improving cluster detection and comparison capabilities[J]. Nucleic Acids Research, 2021.DOI:10.1093/nar/gkab335. |
| [19] | 龙青山. 杂红素生物合成中缩合酶RedH与SDR家族氧化还原酶HbnC的功能与催化机制研究[D]. 上海: 上海交通大学, 2020. |
| LONG Qingshan. Function and catalytic mechanism of condensase RedH and SDR family oxidoreductase HbnC in heterirubin biosynthesis[D]. Shanghai: Shanghai Jiaotong University, 2020. | |
| [20] | Liu Y, Gong R, Liu X, et al. Discovery and characterization of the tubercidin biosynthetic pathway from Streptomyces tubercidicus NBRC 13090[J]. Microbial Cell Factories, 2018, |
| [21] |
McCarty R M, Bandarian V. Biosynthesis of pyrrolopyrimidines[J]. Bioorg Chem, 2012, 43:15-25.
DOI PMID |
| [22] | 张晓敏, 李美宁, 王文卓, 等. 5-碘代杀结核菌素对小鼠神经管的影响及其作用机制[J]. 中国生物制品学杂志, 2021, (6): 682-687. |
| ZHANG Xiaomin, LI Meining, WANG Wenzhuo, et al. Effects of 5-iodicidal tuberculin on the mouse neural tube and its mechanism of action[J]. Chinese Journal of Biological Products, 2021,(6): 682-687. | |
| [23] | 刘启洪, 陈雷, 万维李, 等. Diazepinomicin关键中间体三甲氧基二苯二氮平的合成[J]. 合成化学, 2012, 20(1): 46-48. |
| LIU Qihong, CHEN Lei, WAN Weili, et al. Synthesis of trimethoxydiphenyldiazepine, a key Diazepinomicin intermediate, and diazepine[J]. Synthetic Chemistry, 2012, 20(1): 46-48. | |
| [24] | Jessica-Thandara Gosse, Ghosh Soumya, Sproule Amanda, et al. Whole Genome Sequencing and Metabolomic Study of Cave Streptomyces Isolates ICC1 and ICC4[J]. Frontiers in Microbiology, 2019, 10. |
| [25] | Marnix-H Medema, de Rond Tristan, Moore Bradley-S. Mining genomes to illuminate the specialized chemistry of life[J]. Nature Reviews. Genetics, 2021, 22(9). |
| No related articles found! |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||