

新疆农业科学 ›› 2023, Vol. 60 ›› Issue (5): 1292-1300.DOI: 10.6048/j.issn.1001-4330.2023.05.029
• 微生物·畜牧兽医 • 上一篇
陈开旭( ), 郭翠洁, 杨帆, 任斐儿, 李晓斌, 刘武军(
), 郭翠洁, 杨帆, 任斐儿, 李晓斌, 刘武军( )
)
收稿日期:2022-09-01
出版日期:2023-05-20
发布日期:2023-05-22
通信作者:
刘武军(1966-),女,河南鹿邑人,教授,博士,硕士生/博士生导师研究方向为动物遗传育种,(E-mail) lwj_ws@163.com作者简介:陈开旭(1983-),男,四川三台人,讲师,博士,研究方向为动物遗传育种,(E-mail) chenkaixu@126.com
基金资助:
CHEN Kaixu( ), GUO Cuijie, YANG Fan, REN Feier, LI Xiaobin, LIU Wujun(
), GUO Cuijie, YANG Fan, REN Feier, LI Xiaobin, LIU Wujun( )
)
Received:2022-09-01
Published:2023-05-20
Online:2023-05-22
Supported by:摘要:
【目的】基于全基因组重测序研究新疆细毛羊遗传多样性。【方法】利用新疆细毛羊的全基因组重测序数据和NCBI下载获得的策勒黑羊、巴音布鲁克羊、阿勒泰羊全基因组重测序数据,检测不同绵羊品种的核苷酸多态性和单倍型多态性,评估其遗传多样性现状,计算群体分化指数、观望/观测杂合度,分析遗传多样性,评价不同绵羊品种现有的遗传变异水平。【结果】全基因组重测序获得了97647435个高质量的常染色单核苷酸多态性(SNP)位点和15886270个插入或缺失(Indel);新疆细毛羊的遗传多样性水平显著低于阿勒泰羊,低于巴音布鲁克羊,略高于策勒黑羊;4个绵羊群体遗传多样性顺序:阿勒泰羊> 巴音布鲁克羊> 策勒黑羊> 新疆细毛羊。【结论】选取的4个绵羊群体中,新疆细毛羊的遗传多样性水平相对较低。
中图分类号:
陈开旭, 郭翠洁, 杨帆, 任斐儿, 李晓斌, 刘武军. 基于全基因组重测序分析新疆细毛羊遗传多样性[J]. 新疆农业科学, 2023, 60(5): 1292-1300.
CHEN Kaixu, GUO Cuijie, YANG Fan, REN Feier, LI Xiaobin, LIU Wujun. Genetic diversity analysis of xinjiang sheep with fine wool based on whole-genome Re-sequencing[J]. Xinjiang Agricultural Sciences, 2023, 60(5): 1292-1300.
图1 DNA 样品 1% 琼脂糖凝胶电泳检测结果 注:M 泳道为λ-HindⅢ DNA Marker,1-10泳道为DNA样品
Fig.1 1% agarose gel electrophoresis of DNA samples Note: M line: λ-HindⅢ DNA Marker, 1-10 line: DNA samples
| 样品编号 Sample number | 纯度 OD260/OD280 | 浓度 concentration (ng/μL) |
|---|---|---|
| XFW01 | 1.83 | 81.426 |
| XFW02 | 1.91 | 67.261 |
| XFW03 | 1.88 | 79.537 |
| XFW04 | 1.97 | 94.253 |
| XFW05 | 1.89 | 88.051 |
| XFW06 | 1.89 | 62.306 |
| XFW07 | 2.07 | 105.364 |
| XFW08 | 1.89 | 78.859 |
| XFW09 | 1.88 | 96.857 |
| XFW10 | 1.95 | 79.925 |
表1 新疆细毛羊DNA样品浓度及纯度检测
Tab.1 Detection of DNA concentration and purity of Xinjiang fine wool sheep
| 样品编号 Sample number | 纯度 OD260/OD280 | 浓度 concentration (ng/μL) |
|---|---|---|
| XFW01 | 1.83 | 81.426 |
| XFW02 | 1.91 | 67.261 |
| XFW03 | 1.88 | 79.537 |
| XFW04 | 1.97 | 94.253 |
| XFW05 | 1.89 | 88.051 |
| XFW06 | 1.89 | 62.306 |
| XFW07 | 2.07 | 105.364 |
| XFW08 | 1.89 | 78.859 |
| XFW09 | 1.88 | 96.857 |
| XFW10 | 1.95 | 79.925 |
| 样品编号 Sample number | 比对率 Mapping rate (%) | 全基因组 覆盖度 Coverage (%) | 测序深度 Sequencing depth | Q20 (%) | Q30 (%) |
|---|---|---|---|---|---|
| XFW01 | 99.01 | 97.81 | 8.943 3X | 97.14 | 92.32 |
| XFW02 | 99.46 | 97.89 | 8.167 2X | 97.22 | 92.48 |
| XFW03 | 99.44 | 97.93 | 7.566 7X | 97.12 | 92.22 |
| XFW04 | 99.28 | 97.80 | 7.223 8X | 96.88 | 91.66 |
| XFW05 | 99.32 | 97.90 | 7.484 3X | 97.22 | 92.41 |
| XFW06 | 74.88 | 97.91 | 6.075 4X | 96.81 | 91.62 |
| XFW07 | 99.22 | 97.87 | 7.613 8X | 96.93 | 91.82 |
| XFW08 | 94.48 | 97.93 | 8.478 5X | 96.91 | 91.81 |
| XFW09 | 99.31 | 97.89 | 7.104 6X | 96.73 | 91.47 |
| XFW10 | 99.38 | 97.91 | 8.189 1X | 96.77 | 91.5 |
表2 全基因组测序数据质量统计
Tab.2 Whole genome sequencing data quality statistics
| 样品编号 Sample number | 比对率 Mapping rate (%) | 全基因组 覆盖度 Coverage (%) | 测序深度 Sequencing depth | Q20 (%) | Q30 (%) |
|---|---|---|---|---|---|
| XFW01 | 99.01 | 97.81 | 8.943 3X | 97.14 | 92.32 |
| XFW02 | 99.46 | 97.89 | 8.167 2X | 97.22 | 92.48 |
| XFW03 | 99.44 | 97.93 | 7.566 7X | 97.12 | 92.22 |
| XFW04 | 99.28 | 97.80 | 7.223 8X | 96.88 | 91.66 |
| XFW05 | 99.32 | 97.90 | 7.484 3X | 97.22 | 92.41 |
| XFW06 | 74.88 | 97.91 | 6.075 4X | 96.81 | 91.62 |
| XFW07 | 99.22 | 97.87 | 7.613 8X | 96.93 | 91.82 |
| XFW08 | 94.48 | 97.93 | 8.478 5X | 96.91 | 91.81 |
| XFW09 | 99.31 | 97.89 | 7.104 6X | 96.73 | 91.47 |
| XFW10 | 99.38 | 97.91 | 8.189 1X | 96.77 | 91.5 |
| 单核苷酸多态性SNP | Indel | ||||
|---|---|---|---|---|---|
| Type (alphabetical order) | Count | Percent(%) | Type (alphabetical order) | Count | Percent(%) |
| DOWNSTREAM | 2 232 463 | 2.29 | DOWNSTREAM | 469 219 | 2.95 |
| EXON | 775 956 | 0.79 | EXON | 234 723 | 1.48 |
| INTERGENIC | 56 743 016 | 58.11 | INTERGENIC | 8 705 661 | 54.80 |
| INTRON | 34 744 752 | 35.58 | INTRON | 5 588 596 | 35.18 |
| SPLICE_SITE_ACCEPTOR | 995 | 0.00 | SPLICE_SITE_ACCEPTOR | 11 111 | 0.07 |
| SPLICE_SITE_DONOR | 2108 | 0.00 | SPLICE_SITE_DONOR | 12865 | 0.08 |
| SPLICE_SITE_REGION | 54 513 | 0.06 | SPLICE_SITE_REGION | 15 675 | 0.10 |
| UPSTREAM | 2 350 282 | 2.41 | UPSTREAM | 684 523 | 4.31 |
| UTR_3_PRIME | 511 316 | 0.52 | UTR_3_PRIME | 88 686 | 0.56 |
| UTR_5_PRIME | 232 034 | 0.24 | UTR_5_PRIME | 75 211 | 0.47 |
表3 新疆细毛羊全基因组重测序数据遗传变异鉴定、过滤和注释
Tab.3 The genetic variation information of SNPs and indels in sheep breeds
| 单核苷酸多态性SNP | Indel | ||||
|---|---|---|---|---|---|
| Type (alphabetical order) | Count | Percent(%) | Type (alphabetical order) | Count | Percent(%) |
| DOWNSTREAM | 2 232 463 | 2.29 | DOWNSTREAM | 469 219 | 2.95 |
| EXON | 775 956 | 0.79 | EXON | 234 723 | 1.48 |
| INTERGENIC | 56 743 016 | 58.11 | INTERGENIC | 8 705 661 | 54.80 |
| INTRON | 34 744 752 | 35.58 | INTRON | 5 588 596 | 35.18 |
| SPLICE_SITE_ACCEPTOR | 995 | 0.00 | SPLICE_SITE_ACCEPTOR | 11 111 | 0.07 |
| SPLICE_SITE_DONOR | 2108 | 0.00 | SPLICE_SITE_DONOR | 12865 | 0.08 |
| SPLICE_SITE_REGION | 54 513 | 0.06 | SPLICE_SITE_REGION | 15 675 | 0.10 |
| UPSTREAM | 2 350 282 | 2.41 | UPSTREAM | 684 523 | 4.31 |
| UTR_3_PRIME | 511 316 | 0.52 | UTR_3_PRIME | 88 686 | 0.56 |
| UTR_5_PRIME | 232 034 | 0.24 | UTR_5_PRIME | 75 211 | 0.47 |
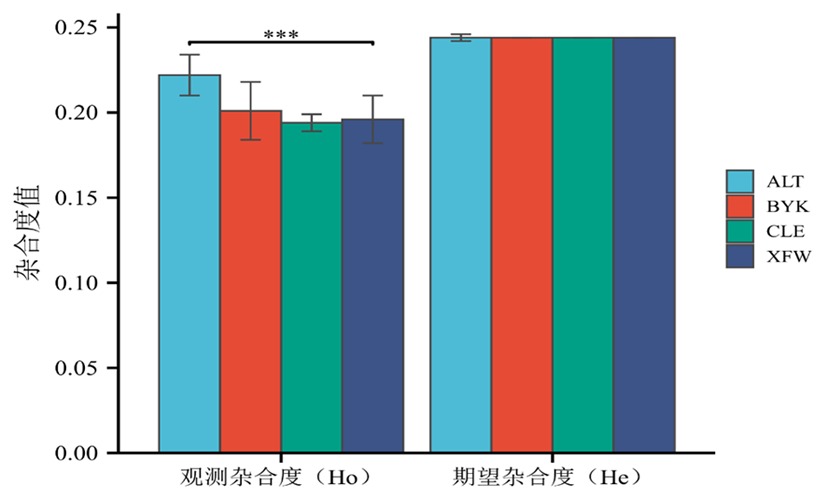
图2 4个绵羊群体的杂合度变化 注:*表示不同绵羊群体间的杂合度具有显著差异(P<0.05),**表示不同绵羊群体间的杂合度具有极显著差异(P<0.01)
Fig.2 Heterozygosity analysis of 4 sheep populations Note:* indicated the heterozygosity of different sheep groups was significantly different (P<0.05),** indicated the heterozygosity of different sheep groups was extremely significantly different
| 品种 Breed | ROH长度 Length of ROH (Mb) | 基因组近交系数 Inbreeding coefficient (FROH) | ||
|---|---|---|---|---|
| 平均值±标准误 Mean±SE | ROH长度区间 Range | 平均值±标准误 Mean±SE | FROH区间 Range | |
| XFW | 110.665±8.744 | 89.609-178.833 | 0.042 3±0.003 2 | 0.034 26-0.068 37 |
| CLE | 93.532±6.956 | 74.687-151.767 | 0.035 8±0.002 5 | 0.028 56-0.058 03 |
| BYK | 88.417±10.694 | 47.985-148.081 | 0.033 8±0.003 9 | 0.018 34-0.056 62 |
| ALT | 74.445±21.225 | 57.063-131.694 | 0.028 5±0.0024 | 0.021 82-0.050 35 |
表4 4个绵羊群体的平均ROH长度和基因组近交系数
Tab.4 Average length and genomic inbreeding coefficient of four sheep groups
| 品种 Breed | ROH长度 Length of ROH (Mb) | 基因组近交系数 Inbreeding coefficient (FROH) | ||
|---|---|---|---|---|
| 平均值±标准误 Mean±SE | ROH长度区间 Range | 平均值±标准误 Mean±SE | FROH区间 Range | |
| XFW | 110.665±8.744 | 89.609-178.833 | 0.042 3±0.003 2 | 0.034 26-0.068 37 |
| CLE | 93.532±6.956 | 74.687-151.767 | 0.035 8±0.002 5 | 0.028 56-0.058 03 |
| BYK | 88.417±10.694 | 47.985-148.081 | 0.033 8±0.003 9 | 0.018 34-0.056 62 |
| ALT | 74.445±21.225 | 57.063-131.694 | 0.028 5±0.0024 | 0.021 82-0.050 35 |
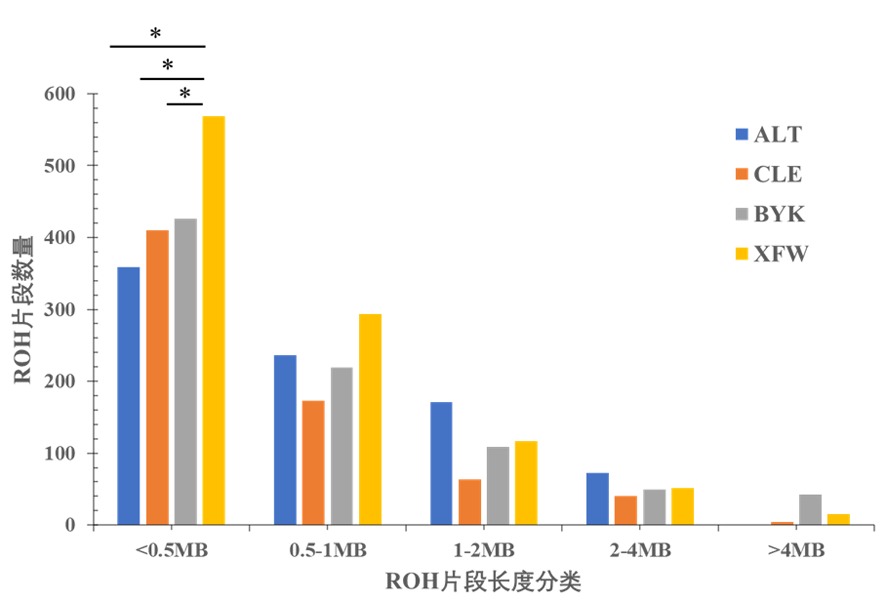
图3 4个绵羊群体的ROH片段长度分类 注:*表示不同绵羊群体间的ROH片段长度差异显著(P<0.05)
Fig.3 The total length of ROH in different length categories Note:* indicated the length of ROH fragment was significantly different among different sheep groups(P<0.05)
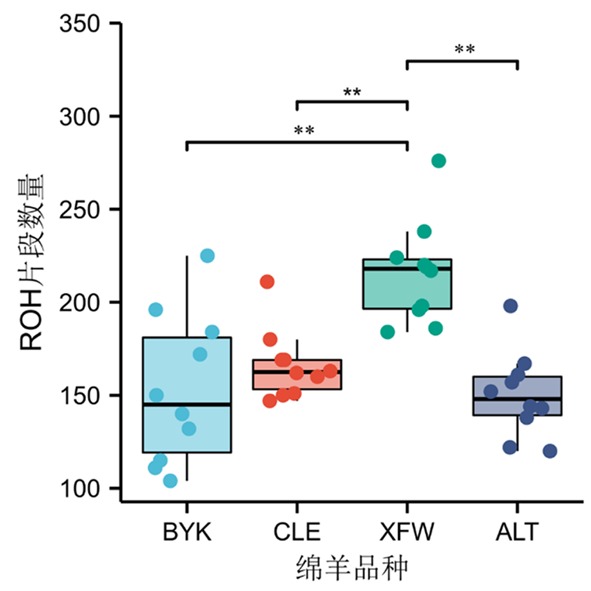
图 4 4个绵羊群体的平均ROH数量和每个个体的 ROH 数量 注:**表示不同绵羊群体间的平均ROH片段数量差异极显著(P<0.01)
Fig.4 The average number of ROH and the number of ROH per individual in 4 sheep groups Note:** indicated the average number of ROH fragments among different sheep groups was extremely significantly different(P<0.01)
| [1] | 刘慧. 中国羊毛贸易发展情况分析[J]. 农业展望, 2010, 6(6): 46-49. |
| LIU Hui. Analysis of the Chinese wool trade development trend[J]. Agricultural Outlook, 2010, 6(6): 46-49. | |
| [2] | 田春英, 王峰, 荣威恒. 内蒙古细毛羊产业可持续发展对策探讨[J]. 畜牧与饲料科学, 2005, 26(4): 48-50. |
| TIAN Chunying, WANG Feng, RONG Weiheng. Countermeasures for sustainable development of Inner Mongolia fine-wool sheep industry[J]. Animal Husbandry and Feed Science, 2005, 26(4): 48-50. | |
| [3] | 赵有璋. 羊生产学[M]. 中国农业出版社, 2011. |
| ZHAO Youzhang. Sheep And Goat Production[M]. China Agriculture Press, 2011. | |
| [4] | 曾献存. 新疆绵羊遗传多样性及主要经济性状候选基因研究[D]. 石河子: 石河子大学, 2010. |
| ZENG Xiancun. Studies on the genetic diversity of Xinjiang sheep breeds and candidate genes of major economic traits[D]. Shehezi: Shihezi University, 2010. | |
| [5] | 沈文, 孙延鸣, 崔茹鹏, 等. 新疆细毛羊KAP6.1基因的克隆及序列分析[J]. 黑龙江畜牧兽医, 2013, 3: 31-34. |
| SHEN Wen, SUN Yanming, CUI Rupeng, et al. Cloning and sequence analysis of KAP 6.1 gene in Xinjiang fine-wool sheep[J]. Heilongjiang Animal Science and Veterinary Medicine, 2013(3): 31-34. | |
| [6] | 肖非. 新疆绵羊种质资源调查、保护及遗传多样性[D]. 石河子: 石河子大学, 2009. |
| XIAO Fei. Investigation, conservation and genetic diversity of germplasm resources in Xinjiang sheep[D]. Shiehzi: Shihezi University, 2009. | |
| [7] | 赵永欣, 李孟华. 中国绵羊起源, 进化和遗传多样性研究进展[J]. 遗传, 2017, 39(11):958-973. |
| ZHAO Yongxin, LI Menghua. Research advances on the origin, evolution and genetic diversity of Chinese native sheep breeds[J]. Hereditas, 2017, 39(11): 958-973. | |
| [8] | 王秀蓉. 基于SSR标记评估青海省家牦牛和藏绵羊遗传多样性及近交程度[D]. 西宁: 青海师范大学, 2019. |
| WANG Xiurong. Study on genetic diversity and inbreeding degree of domestic Yak and Tibetan sheep in Qinghai Province based on SSR markers [D]. Xining: Qinghai Normal University, 2019. | |
| [9] | Li H. Aligning sequence reads, clone sequences and assembly contigs with BWA-MEM[J]. arXiv e-prints, 2013: arXiv: 1303.3997. |
| [10] |
Li H, Handsaker B, Wysoker A, et al. The sequence alignment/map format and SAM tools[J]. Bioinformatics, 2009, 25(16): 2078-2079.
DOI URL |
| [11] |
McKenna A, Hanna M, Banks E, et al. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data[J]. Genome Research, 2010, 20(9): 1297-1303.
DOI PMID |
| [12] | Wang K, Li M, Hakonarson H. ANNOVAR: functional annotation of genetic variants from high-throughput sequencing data[J]. Nucleic Acids Research, 2010, 38(16): e164. |
| [13] |
Yang H, Wang K. Genomic variant annotation and prioritization with ANNOVAR and ANNOVAR[J]. Nature Protocols, 2015, 10(10): 1556-1566.
DOI PMID |
| [14] |
Haubold B, Pfaffelhuber P, Lynch M. mlRho-a program for estimating the population mutation and recombination rates from shotgun-sequenced diploid genomes[J]. Molecular Ecology, 2010, 19: 277-284.
DOI URL |
| [15] |
Purcell S, Neale B, Todd-Brown K, et al. PLINK: a tool set for whole-genome association and population-based linkage analyses[J]. The American Journal of Human Genetics, 2007, 81(3): 559-575.
DOI URL |
| [16] | Bosse M, Megens H J, Madsen O, et al. Regions of homozygosity in the porcine genome: consequence of demography and the recombination landscape[J]. PLoS Genetics, 2012, 8(11): e1003100. |
| [17] |
Ceballos F C, Joshi P K, Clark D W, et al. Runs of homozygosity: windows into population history and trait architecture[J]. Nature Reviews Genetics, 2018, 19(4): 220.
DOI PMID |
| [18] |
Garvin M R, Saitoh K, Gharrett A J. Application of single nucleotide polymorphisms to non‐model species: a technical review[J]. Molecular Ecology Resources, 2010, 10(6): 915-934.
DOI PMID |
| [19] | Fan B, Du Z Q, Gorbach D M, et al. Development and application of high-density SNP arrays in genomic studies of domestic animals[J]. Asian-Australasian Journal of Animal Sciences, 2010, 23(7): 833-847. |
| [20] | 张彦斌, 罗海涛, 范鑫, 等. 锡林郭勒绵羊品种遗传多样性研究进展[J]. 中国畜禽种业, 2020, 16(3): 3-6. |
| ZHANG Yanbin, LUO Haitao, FAN Xin, et al. Research progresses of the varietal genetic polymorphism of Xilinguole sheep[J]. The Chinese Livestock and Poultry Breeding, 2020, 16(3): 3-6. | |
| [21] |
Ceballos F C, Joshi P K, Clark D W, et al. Runs of homozygosity: windows into population history and trait architecture[J]. Nature Reviews Genetics, 2018, 19(4): 220.
DOI PMID |
| [22] |
Broman K W, Weber J L. Long homozygous chromosomal segments in reference families from the centre d'Etude du polymorphisme humain[J]. The American Journal of Human Genetics, 1999, 65(6): 1493-1500.
DOI URL |
| [23] |
Caballero A, Villanueva B, Druet T. On the estimation of inbreeding depression using different measures of inbreeding from molecular markers[J]. Evolutionary Applications, 2021, 14(2): 416-428.
DOI PMID |
| [24] |
D’ambrosio J, Phocas F, Haffray P, et al. Genome-wide estimates of genetic diversity, inbreeding and effective size of experimental and commercial rainbow trout lines undergoing selective breeding[J]. Genetics Selection Evolution, 2019, 51(1): 1-15.
DOI |
| [25] |
Letko A, Minor K M, Jagannathan V, et al. Genomic diversity and population structure of the Leonberger dog breed[J]. Genetics Selection Evolution, 2020, 52(1): 1-12.
DOI |
| [26] | Fang Y, Hao X, Xu Z, et al. Genome-wide detection of runs of homozygosity in Laiwu pigs revealed by sequencing data[J]. Frontiers in Genetics, 2021: 463. |
| [27] |
Liu D, Chen Z, Zhao W, et al. Genome-wide selection signatures detection in Shanghai Holstein cattle population identified genes related to adaption, health and reproduction traits[J]. BMC Genomics, 2021, 22(1): 1-19.
DOI |
| [28] |
Gorssen W, Meyermans R, Janssens S, et al. A publicly available repository of ROH islands reveals signatures of selection in different livestock and pet species[J]. Genetics Selection Evolution, 2021, 53(1): 1-10.
DOI |
| [29] |
Stoffel M A, Johnston S E, Pilkington J G, et al. Mutation load decreases with haplotype age in wild Soay sheep[J]. Evolution Letters, 2021, 5(3): 187-195.
DOI PMID |
| [30] |
Zhang X, Kim B, Lohmueller K E, et al. The impact of recessive deleterious variation on signals of adaptive introgression in human populations[J]. Genetics, 2020, 215(3): 799-812.
DOI PMID |
| [31] | Khan A, Patel K, Shukla H, et al. Genomic evidence for inbreeding depression and purging of deleterious genetic variation in Indian tigers[J]. Proceedings of the National Academy of Sciences, 2021, 118(49): e2023018118. |
| [32] | Addo S, Klingel S, Thaller G, et al. Genetic diversity and the application of runs of homozygosity-based methods for inbreeding estimation in German White-headed Mutton sheep[J]. PloS One, 2021, 16(5): e0250608. |
| [33] | Suezawa R, Nikadori H, Sasaki S. Genetic diversity and genomic inbreeding in Japanese Black cows in the islands of Okinawa Prefecture evaluated using single‐nucleotide polymorphism array[J]. Animal Science Journal, 2021, 92(1): e13525. |
| [34] | Roh H J, Kim S C, Cho C Y, et al. Estimating genetic diversity and population structure of 22 chicken breeds in Asia using microsatellite markers[J]. Asian-Australasian Journal of Animal Sciences, 2020, 33(12): 1896. |
| [35] | Humble E, Paijmans A J, Forcada J, et al. An 85K SNP array uncovers inbreeding and cryptic relatedness in an Antarctic fur seal breeding colony[J]. G3: Genes, Genomes, Genetics, 2020, 10(8): 2787-2799. |
| [36] | 杨湛澄, 黄河天, 闫青霞, 等. 利用高密度 SNP 标记分析中国荷斯坦牛基因组近交[J]. 遗传, 2017, 39(1): 16-23. |
| YANG Zhancheng, HUANG Hetian, YAN Qingxia, et al. Estimation of genomic inbreeding coefficients based on high-density SNP markers in Chinese Holstein cattle[J]. Hereditas, 2017, 39(1): 16-23. | |
| [37] |
Stoffel M A, Johnston S E, Pilkington J G, et al. Genetic architecture and lifetime dynamics of inbreeding depression in a wild mammal[J]. Nature Communications, 2021, 12(1): 1-10.
DOI |
| [38] |
Ghoreishifar S M, Moradi-Shahrbabak H, Fallahi M H, et al. Genomic measures of inbreeding coefficients and genome-wide scan for runs of homozygosity islands in Iranian river buffalo, Bubalus bubalis[J]. BMC Genetics, 2020, 21(1): 1-12.
DOI |
| [39] | Suezawa R, Nikadori H, Sasaki S. Genetic diversity and genomic inbreeding in Japanese Black cows in the islands of Okinawa Prefecture evaluated using single‐nucleotide polymorphism array[J]. Animal Science Journal, 2021, 92(1): e13525. |
| [40] | Lewontin R C, Kojima K. The evolutionary dynamics of complex polymorphisms[J]. Evolution, 1960: 458-472. |
| [41] |
Slatkin M. Linkage disequilibrium—understanding the evolutionary past and mapping the medical future[J]. Nature Reviews Genetics, 2008, 9(6): 477-485.
DOI |
| [1] | 曾婉盈, 耿洪伟, 程宇坤, 李思忠, 钱松廷, 高卫时, 张立明. 甜菜品系叶丛快速生长期抗旱性综合评价[J]. 新疆农业科学, 2024, 61(9): 2140-2151. |
| [2] | 徐毛毛, 高杰, 李君明, 李鑫, 刘磊, 潘峰. 20个番茄商业品种群体的多样性分析[J]. 新疆农业科学, 2024, 61(9): 2191-2196. |
| [3] | 苗雨, 陈翠霞, 马艳明, 邢国芳, 董裕生, 陈智军, 王仙, 向莉. 276份中亚大麦种质资源表型性状的遗传多样性分析[J]. 新疆农业科学, 2024, 61(8): 1888-1895. |
| [4] | 杨璐, 王娜, 范少丽, 程平, 李宏, 汪阳东. 黑桑种质资源表型性状变异特征分析[J]. 新疆农业科学, 2024, 61(5): 1172-1181. |
| [5] | 王帆, 李玉姗, 王威, 邓超宏, 赵连佳, 马越, 肖菁, 庄红梅, 许红军. 基于表型性状鉴定及SSR分子标记分析芜菁种质资源遗传多样性[J]. 新疆农业科学, 2024, 61(11): 2601-2613. |
| [6] | 李超, 杨英, 郑贺云, 杨建丽, 陈伟, 杨咪, 孙玉萍. 基于SSR荧光标记分析新疆甜瓜种质资源遗传多样性与群体结构[J]. 新疆农业科学, 2024, 61(11): 2614-2625. |
| [7] | 汪天玲, 侯献飞, 施俊杰, 孙全喜, 贾东海, 顾元国, 单世华, 苗昊翠, 李强. 67份匍匐型花生种质资源遗传多样性分析[J]. 新疆农业科学, 2024, 61(1): 42-54. |
| [8] | 逯涛, 曾庆涛, 张文, 王文博, 王政洋, 杨芮, 孙玉岩. 主成分分析及灰色关联度分析综合评价棉花产量与品质[J]. 新疆农业科学, 2023, 60(5): 1099-1109. |
| [9] | 杨建超, 徐诚, 杨平, 杨鸿基, 张凯浩, 马新超, 高亚宁, 陈有福, 付湘, 轩正英. 氮素指数施肥对设施薄皮甜瓜养分承载及氮吸收利用的影响[J]. 新疆农业科学, 2023, 60(3): 590-600. |
| [10] | 罗影, 努尔孜亚·亚力买买提, 贾文捷, 刘洁莹, 贾培松. 新疆野生大环柄菇的分子鉴定与遗传多样性分析[J]. 新疆农业科学, 2023, 60(10): 2501-2508. |
| [11] | 姚莹莹, 李家辉, 李海英, 吴盈萍, 赵晓钰, 周军, 赵全庄, 李宗福. 也迷里鸡微卫星遗传多样性分析[J]. 新疆农业科学, 2023, 60(10): 2574-2582. |
| [12] | 赵双印, 王为然, 闫雪雪, 比买热木·阿不都艾海提, 董洁, 吐尔逊·吐尔洪, 阿里甫·艾尔西. 120份棉花种质资源品质性状与遗传多样性分析[J]. 新疆农业科学, 2023, 60(1): 17-24. |
| [13] | 李岳雁, 李玉姗, 王帆, 郭雅文, 王飞燕, 高杰, 宋羽. 不同品种番茄果实性状遗传多样性及聚类分析[J]. 新疆农业科学, 2022, 59(9): 2147-2157. |
| [14] | 袁雷, 吉雪花, 张国儒, 石林媛, 郭河瑶, 唐亚萍, 杨涛, 杨生保. 52份辣椒主要果实性状的遗传多样性及聚类分析[J]. 新疆农业科学, 2022, 59(8): 1935-1944. |
| [15] | 吴巧玉, 何天久. 紫色甘薯资源的形态学和ISSR荧光标记分析[J]. 新疆农业科学, 2022, 59(7): 1625-1631. |
| 阅读次数 | ||||||||||||||||||||||||||||||||||||||||||||||||||
|
全文 75
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||
|
摘要 330
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||